Editor’s Note
This article was originally published 26 November 2020. We have updated it to include additional comments from independent experts.
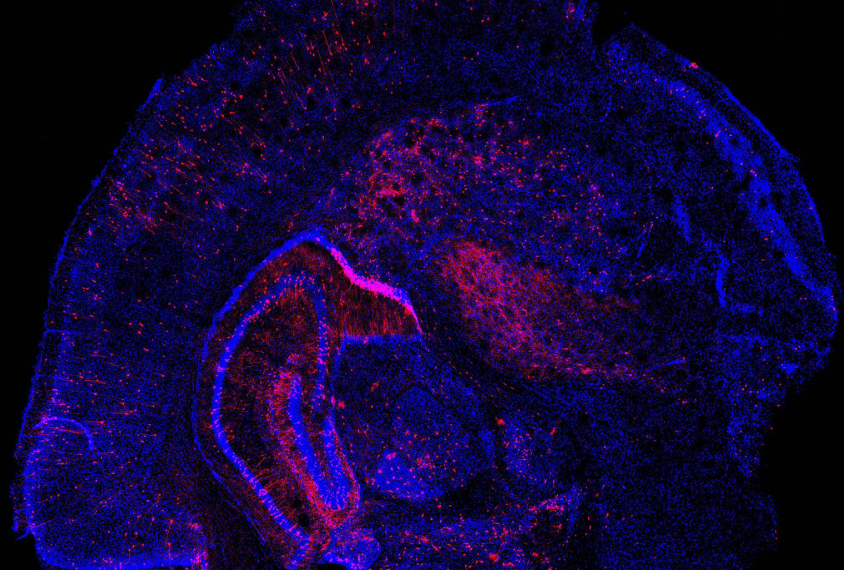
Researchers have devised a new way of seeing how gene mutations affect cells during development.
This article was originally published 26 November 2020. We have updated it to include additional comments from independent experts.
Mutations in a set of 35 genes linked to autism affect both neurons and glial cells in developing mice, according to a study published in November in Science. The study is the first to screen multiple autism-gene mutations, one by one, in living mice and analyze their effects in individual brain cells.
The findings support the idea that mutations in autism genes reverberate through multiple cell types, reflecting autism’s complexity, says study investigator Xin Jin, a junior fellow at Harvard University. That’s not surprising, she says, considering that many autism genes encode proteins called transcription factors, which regulate the activity of many other genes.
“These are general molecules affecting development,” Jin says.
About 100 genes have been strongly linked to autism, and sequencing efforts continue to uncover more. But how mutations in each of these genes affect brain development remains largely unknown. Although researchers have investigated the molecular effects of just a few genes, testing all of them individually is a daunting task.
In the new work, researchers built on a technique called Perturb-seq, developed in 2016. This technique mutates the genome in individual cells using CRISPR technology and then sequences the RNA in each cell to determine how the mutation changed it.
Perturb-seq has been used in cells grown in a dish, says Paola Arlotta, professor of stem cell and regenerative biology at Harvard University, who co-led the work, “but this is the first time that it is applied in an intact, living and developing organism.”
The study “is a proof of principle that this method can be used successfully to generate data for a large pool of genes at once,” says Donna Werling, assistant professor of genetics at the University of Wisconsin-Madison, who was not involved in the work.
Jin, Arlotta and their colleagues injected viruses carrying CRISPR gene-editing machinery into 12-day-old mouse embryos. They used just enough virus to cause mutations in 1 out of every 1,000 stem cells in the embryonic brain. The 35 genes they mutated were identified as autism genes in a January study.
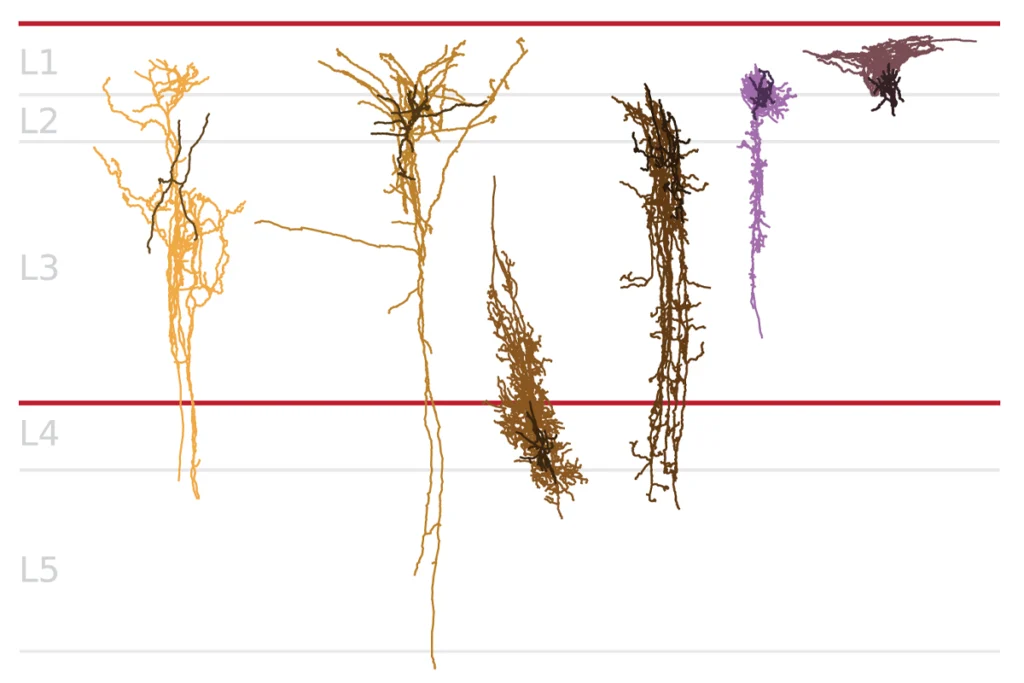
When the pups were 7 days old, the researchers sequenced the RNA in five different cell types in the pups’ brains: three types of neurons and two types of glial cells. Each cell they sequenced carried just one mutation, allowing them to assess that mutation’s effect on each of the five cell types.
By applying machine learning to gene-expression data from unmutated cells of each type, they identified a total of 14 gene modules — sets of genes involved in related biological processes . Mutations in 9 of the 35 autism-linked genes affect five modules in multiple cell types, they found.
The approach enables researchers to study how different autism-linked genes might converge on similar developmental pathways, Jin says.
“This study is a beautiful example of how [by] combining CRISPR-Cas9 with single-cell gene-expression profiling we can begin to disentangle the complexity of genetically heterogenous disorders such as [autism],” says Gaia Novarino, professor of neuroscience at the Institute of Science and Technology in Klosterneuburg, Austria.
Conducting similar studies at different time points during development might reveal how the effects of different mutations change over time, she says.
Of course, people who have mutations generally carry them in all their cells, Werling says, so creating them in a small selection of cells throughout the brain doesn’t reflect their impact on development. But the trade-off is being able to study multiple genes at once.
“This is super-exciting work, and I think this represents a starting point for a whole array of applications for this sort of technology,” Werling says.
The researchers plan to expand the number of genes that the technique can mutate at once, Arlotta says. They also plan to apply it to lab-grown human brain cells that form clusters called organoids.
“That would allow us to really understand whether these genes are affecting the development of a human brain differently than the development of a mouse brain,” she says.