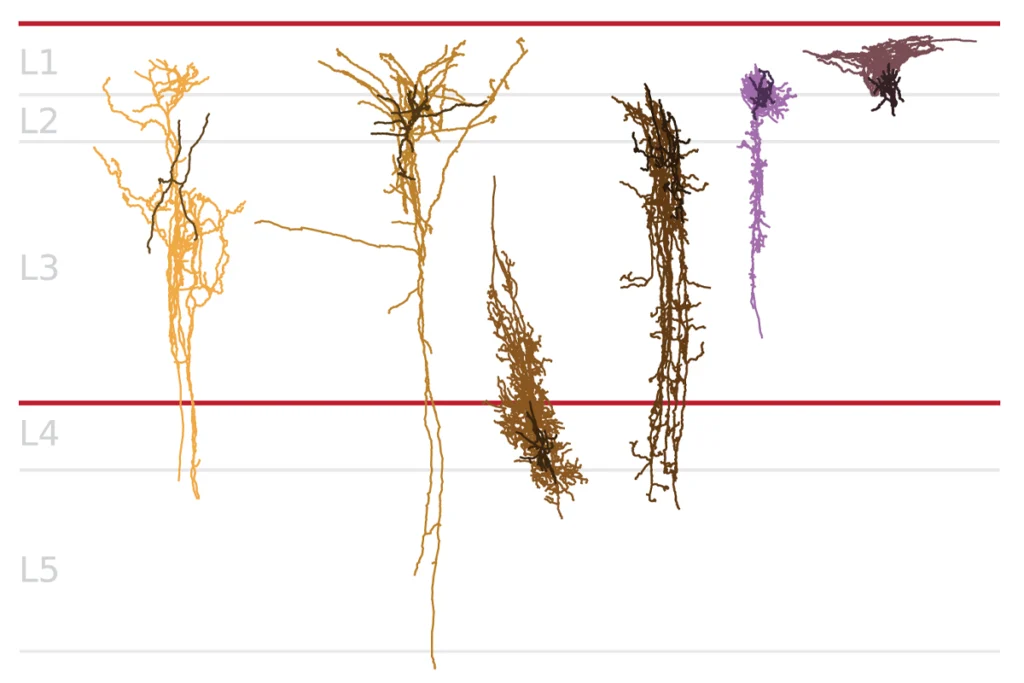
Methylation map of neurons points to autism-related genes
Large-scale patterns of methylation throughout the genome affect the expression of many autism-related genes and may contribute to the disorder’s development, according to a study published online Friday in Genome Research. The study is among the first to look at genome-wide patterns of methylation in brain cells.

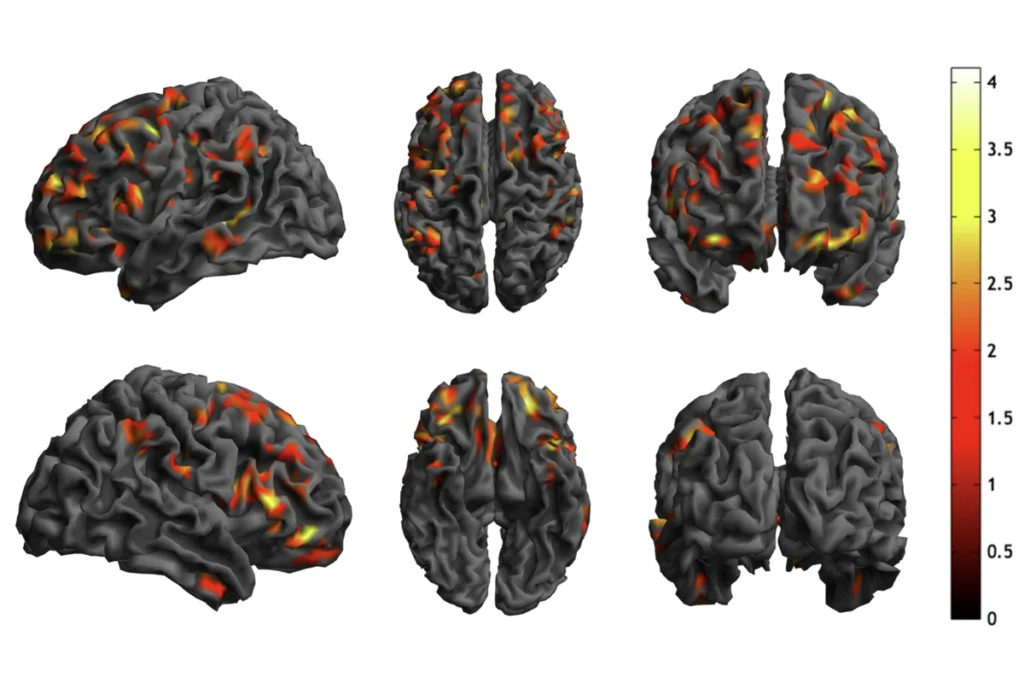
Rainbow bright: Different levels of methylation across the genome indicate areas rich in autism-related genes.
Large-scale patterns of methylation throughout the genome affect the expression of many autism-related genes and may contribute to the disorder’s development, according to a study published online Friday in Genome Research1.
Methylation is one type of epigenetic change, a chemical alteration to DNA that affects its function but not its genetic code. The new study is among the first to take advantage of new methods to look at genome-wide patterns of methylation in brain cells.
“Now we can see a big picture of where methylation is in the genome,” says lead investigator Janine LaSalle, professor of medical microbiology and immunology at the University of California, Davis. She compares the new technology to “satellite images looking down at the whole landscape of the epigenome.”
This bird’s-eye view highlights regions containing many autism-related genes that are also affected by methylation, the new study found. “This really adds a new dimension of complexity to our understanding of how methylation may be regulated in the brain and in autism,” says Valerie Hu, professor of biochemistry and molecular biology at George Washington University in Washington, DC, who was not involved in the work.
Scientists have long known that methylation, or the addition of a methyl group to certain DNA bases, helps determine which genes are turned on and off in different cells. But previous studies have mostly focused on specific, relatively small parts of the genome — especially promoters, which act as on-off switches for genes.
In the past few years, the development of rapid, high-throughput DNA sequencing technology has enabled researchers to look at methylation more broadly.
These methods have shown that most of the genome is rich in methyl groups, but there are some areas, tens of thousands of base pairs long, that have lower levels of methylation. Genes in these less-methylated regions are also expressed at lower levels. That finding contrasts with the conventional understanding of methylation: At promoters, for example, methylation generally turns genes off.
“It’s a level of regulation that wasn’t understood until recently,” says Joseph Ecker, director of the Genomic Analysis Laboratory at the Salk Institute for Biological Studies in San Diego, California. Ecker led the effort to make the first high-resolution human methylation map in 20092.
Form and function:
In the new study, LaSalle and her colleagues compared methylation patterns of skin cells and neurons. They found that some parts of the genome have high levels of methylation in neurons, but more modest levels in skin cells. These regions, which the researchers dubbed neuronal highly methylated domains, or HMDs, contain about 2,950 genes that are more active in neurons than in skin cells.
When the researchers looked up the function of these genes, “it just lit up the synapse,” the junction between neurons, LaSalle says. Many of the genes code for proteins found at the connections between neurons and needed for sending signals between them. About 60 of the genes, a significantly larger number than would be expected by chance, have also been linked to autism.
But any long list of genes active in neurons is bound to turn up a few that are implicated in autism, notes Hongjun Song, professor of neurology at Johns Hopkins University School of Medicine in Baltimore. “I don’t know how specifically the data is linked to autism versus any other neurological disorders.”
LaSalle acknowledges the criticism, and notes that in a separate unpublished analysis, her team found that the regions are also enriched in genes associated with some mental illnesses such as anxiety and depression, but not other disorders, including Alzheimer’s disease.
“We wouldn’t claim that these genes are uniquely enriched for autism candidate genes, but they do seem to be somewhat specific to neurodevelopmental and mental health disorders,” LaSalle says.
The neurons used in the study, called SH-SY5Y cells, are cancer cells that have been growing in the laboratory for many years — both factors that can affect methylation patterns, Song points out. “We don’t know how much this reflects the physiological cell type in human brains,” he says. “But this is a good start. We now have a reference point to look at.”
An important next step would be to determine whether neurons from people with autism have different methylation patterns than do neurons from controls.
Some methylation differences have already been identified in people with autism. For example, identical twin pairs in which one twin has autism and the other does not show specific autism-linked patterns of methylation at certain genomic sites called CpG islands3.
Methylation patterns have the potential to illuminate how environmental factors such as diet and exposure to toxins may affect autism risk4, says LaSalle.
For example, a study published last month reported that when women take prenatal vitamins prior to and immediately after conception, their children have a lower risk of autism5. Prenatal vitamins contain large amounts of folic acid, a rich source of methyl groups.
LaSalle says her group’s new study provides some hints about which autism-related genes may be affected by such environmental factors. “It’s basically giving us an epigenetic road map of where to look for DNA methylation alterations in autism in the future.”
References:
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question