Random genetic changes may explain variability in autism
Random changes in gene expression can cause genetically identical embryos to develop different traits, according to a study of worms published in Nature. The findings suggest that haphazard movements of molecules could partly explain why autism-associated mutations don’t always cause the same symptoms.
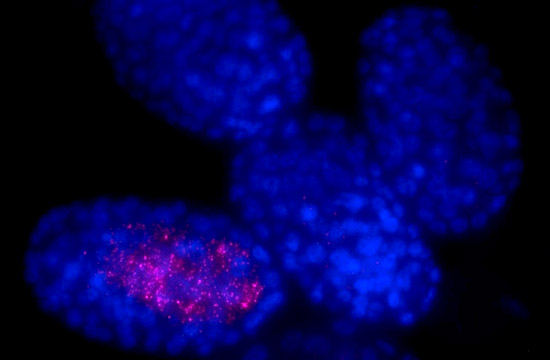
Chance occurrence: These worm embryos carry the same DNA, but only one produces the molecules (pink) needed to make gut cells.
Random changes in gene expression can cause genetically identical embryos to develop different traits, according to a study of worms published 18 February in Nature1.
The findings suggest that haphazard movements of molecules could partly explain why identical twins can look and behave differently, and why autism-associated genetic mutations don’t always cause the same behavioral effects, researchers say.
“This elegant model in genetically identical worms provides some of the first evidence that random differences in gene expression may explain some of the variability we observe in symptoms of autism,” says Donna Martin, professor of pediatrics and human genetics at the University of Michigan, who was not involved in the new study.
Gene expression — the process of converting DNA to protein — occurs when specific genes or chemicals turn a gene ‘on’. Expression is influenced by any number of genetic and environmental factors, from feelings of loneliness2 to chemical exposures3. The new study is the first to show that random changes in the cellular environment can also dictate whether a gene is switched on or off.
Cellular processes that regulate gene expression usually operate without a hitch. But because molecules are constantly bouncing around inside a cell, they occasionally interact in random ways. In these aberrant cases, DNA may fail to switch on a gene that’s normally active, the study found.
The researchers investigated the development of intestinal cells in genetically identical worms, Caenorhabditis elegans. Unlike humans, worms can be grown in identical environments. The embryos are also translucent, making their DNA easy to visualize.
The scientists first characterized the normal progression of intestine development in C. elegans using a fluorescent tag that binds to messenger RNA — molecules that carry instructions for building proteins.
The tag allowed the researchers to visualize and count the number of copies of a particular messenger RNA sequence; this number indicates whether a new gene will be turned on.
The development of the 20 cells that make up the normal adult worm’s intestine follows a simple regulatory network. The most important gene in the cascade, called SKN-1, activates a series of four other genes. Ultimately, this turns on a final gene, ELT-2, which prompts embryonic cells to differentiate into gut cells.
When scientists introduce a mutation in SKN-1, however, some worms develop normal digestive tracts and others do not. In mutant embryos, SKN-1 does not express three genes in the pathway, leaving only one gene, END-1, to activate ELT-2. Although the researchers don’t yet know why, END-1’s expression varies significantly among the embryos, ranging from extremely low to nearly normal levels.
On closer examination, the researchers found that the mutant embryos that produce low levels of END-1 messenger RNA never express ELT-2, and never develop a digestive tract. Conversely, those that have normal levels of END-1 do express ELT-2 and develop normal digestive tracts.
“The idea that organisms require a threshold of gene activity to manifest a specific trait likely also explains random genetic variation we see in humans with genetic diseases like autism,” says lead investigator Scott Rifkin, professor of biophysics at the Massachusetts Institute of Technology (MIT).
Outside the Petri dish:
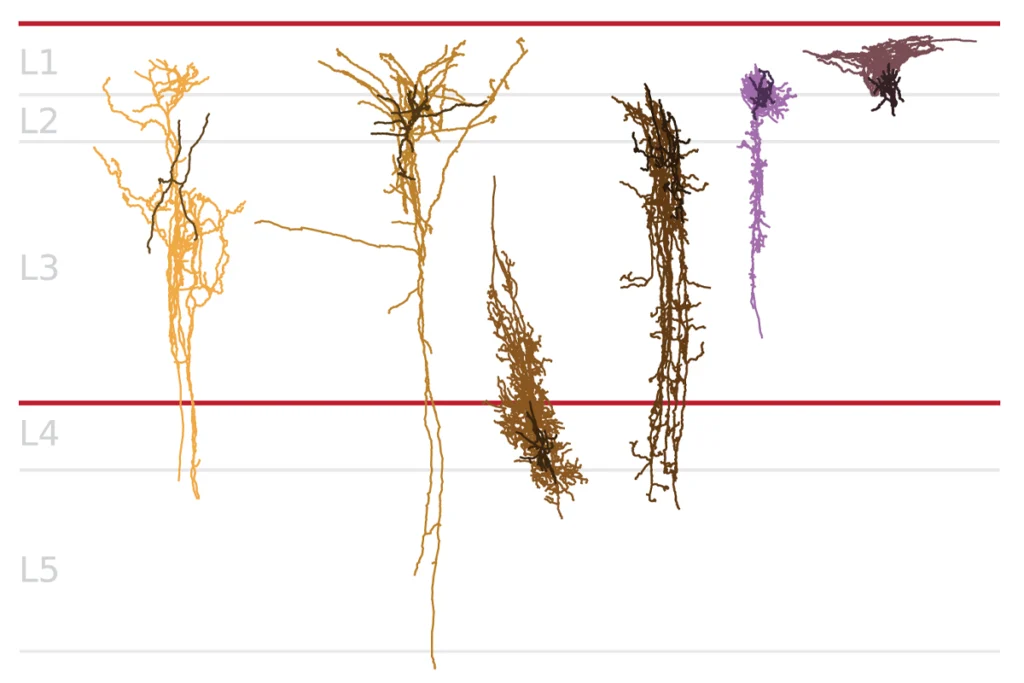
Just as SKN-1 regulates the expression of genes that form gut cells in worms, some autism-related genes regulate pathways important for brain development.
“We are still learning how relevant certain genes are for autism,” Martin says. “If mutations on these genes are associated with different aspects of the disorder, it’s clear that they do not cause the same characteristics in every individual.”
For instance, a study published last year showed that variations in the MeCP2 gene — which is mutated in Rett syndrome, an autism spectrum disorder — can disrupt brain development and lead to symptoms of autism in boys4. In girls with the same mutation, however, the severity of symptoms varies, indicating that the mutation does not affect everyone to the same degree.
Another gene, HOXA1, which helps govern an embryo’s head and brainstem development, may also regulate genes related to autism. Researchers at the University of Rochester found that 39 percent of people with autism carry a mutation in one of the two copies of HOXA15. Not everyone with the mutation exhibits symptoms of the disorder, however.
But studying these random variations in people is not so simple. For instance, because RNA degrades rapidly, gene expression studies require fresh tissue.
“It’s hard to get fresh brain tissue, especially from live patients, so the most accessible choice is blood, which may not reflect the same expression pattern as the brain,” says David Ledbetter, professor of human genetics at Emory University.
Human development is also much more complex — and much less easily manipulated — than that of worms, Rifkin adds. It is virtually impossible, for instance, to isolate people in identical environments, but you can place worms in identical Petri dishes to eliminate external confounders.
Still, the processes involved in DNA expression are very similar between the two species, says Rifkin, and researchers should explore random gene expression in more complex organisms.
References:
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question