Should we use the computational or the network approach to analyze functional brain-imaging data—why not both?
Emerging methods make it possible to combine the two tactics from opposite ends of the analytic spectrum, enabling scientists to have their cake and eat it too.
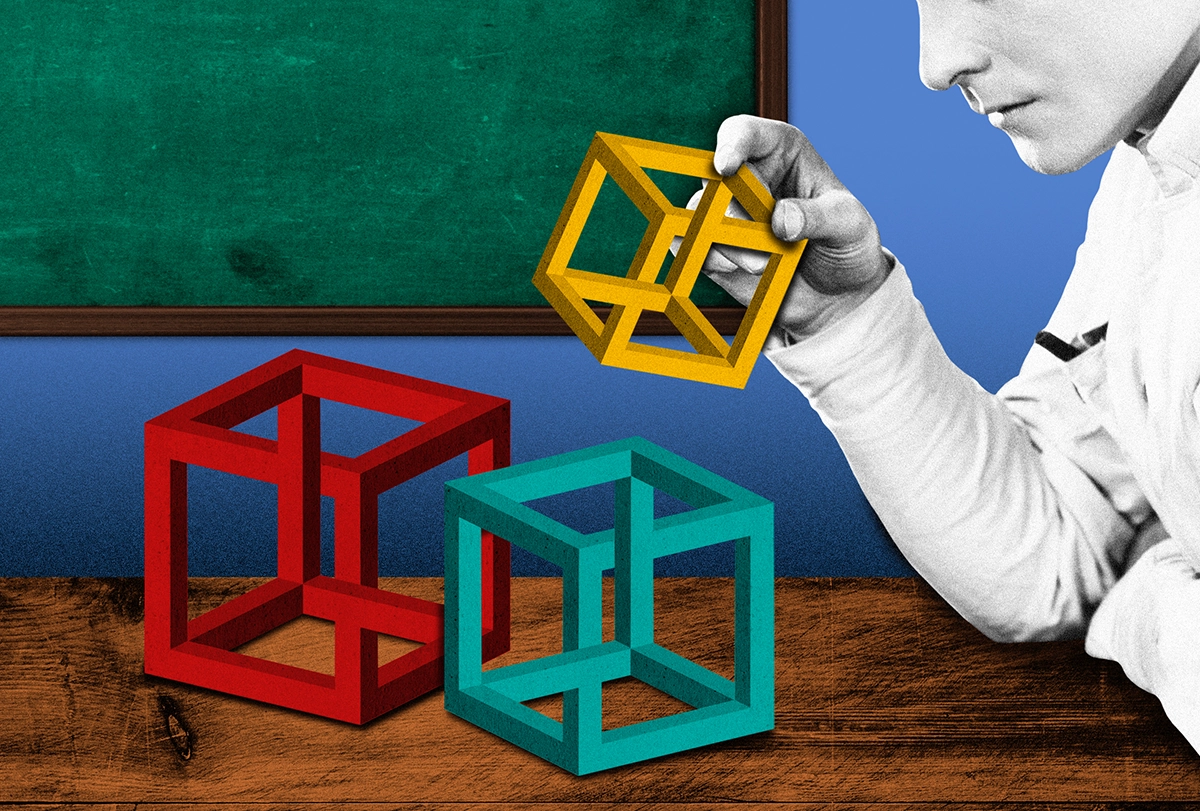
Imagine the challenge for a modern neuroscientist interested in something seemingly trivial, such as how someone can interpret a bistable image in two different ways. A well-known example is the Necker cube (see image below): With a switch of perspective, you can see the cube as projecting into or out of the page. (If you’re struggling to see these differences, view the shaded blue panels on the right as if they are in the foreground and then look back at the translucent cube.)
Once you know the trick, you can make the switch in perspective voluntarily, but how does your brain achieve this seemingly trivial feat? In other words, how do coordinated interactions among the billions of neurons in our brain give rise to a change in our interpretation of an ambiguous image?
For the sake of the example, let’s spin a little further: Imagine that our neuroscientist has access to whole-brain functional neuroimaging data (such as fMRI) collected while a group of healthy participants performed a perceptual task just like this one in an MRI scanner. How can these data help her to understand how the brain accomplished the task?
Although there are now countless ways to analyze functional neuroimaging data, two main approaches feature in the literature—the brain/network mapping approach and the computational approach. By combining these approaches, as I will explain below, our hypothetical neuroscientist might reach a more nuanced understanding of how bistable perception works.
W
ith the brain/network mapping approach, the idea is to analyze neuroimaging data using statistical techniques that can summarize patterns inherent within the data and then ask whether the different aspects of the task are associated with different network signatures. The classical approach is to treat each reconstructed region of the brain signal as an independent sample and determine which voxels’—3D pixels’—activity correlated most closely in time with changes in the task.This tactic has led to many insights into brain regions and their functional signatures. Take, for instance, the pioneering work of New York University neuroscientist David Heeger and his colleagues, which demonstrated blood-flow changes in the visual cortex that covaried with the dominant visual stimulus in a rivalry paradigm like the Necker cube example above.
This mapping approach identifies the areas of the brain involved, but it doesn’t get us much closer to the underlying mechanism. To understand how individual brain regions work together to fulfill more complex functions, such as conscious perception, researchers can instead analyze data as part of an interconnected network—a graph with nodes interconnected by a set of edges—and leverage the insights of graph theory, a powerful suite of mathematical tools.
To do so, scientists must first define the nodes and edges of a network, as well as choose which family of subsequent measures to apply to the graph. Let’s say we choose a pre-defined set of regions in the ventral temporal lobe, thalamus and superior colliculus as nodes, and the temporal correlations between their activity patterns as the edges. Graph theory enables us to track higher-order structure in the correlation patterns as a function of our flickering perceptual interpretation of the Necker cube. Studies that use such approaches have shown that much larger networks—extending outside the specialized visual circuits of the brain—are involved in visual rivalry.
An alternative strategy is the computational approach. This method starts by making assumptions about how a participant solves a task. In the case of the Necker cube, for example, we might assume that switching occurs according to a white-noise process that evolves toward to two fixed-point attractors. Or we might assume that the problem is solved by a hidden layer within a recurrent neural network (RNN) that learns to accumulate evidence for different interpretations of the stimuli.
The white-noise assumptions rely on particular classes of mathematical functions. Though often elegant, they may not represent the mechanism the brain uses to solve the task, but rather an abstraction that can help us understand the potential computations at play. An illustrative example comes from York University neuroscientist Hugh Wilson, who was able to create a minimalistic network of mathematical equations that could reproduce rivalry-like behavior. The RNN assumptions are instead focused on whether the simplified model trained on the task—for instance, an RNN with distinctly non-brain-like features such as identical nodes that use back-propagation to train their connection weights—is a decent simulation of the brain. (We recently used this approach to understand how the brain processes visual ambiguity). The experimenter then interrogates their data for signatures associated with these computational building blocks, often using statistical techniques to ensure that the match between model and data is more profound than one that could arise simply as a result of noise.
S
o what are we to conclude from these different approaches? When we use computational approaches, we retain control over the interpretability of our conclusions, but at the risk of a mismatch between our assumptions and the as-yet-unknowable ground truth of the functional organization of the brain. By contrast, when we use brain/network mapping approaches, we hopefully provide a better match to the brain as it is, but neural activity is highly dynamic and multiscale, and static maps of the brain simply can’t match this level of complexity. This means that we struggle to shift between static maps of brain activity and the dynamic neural mechanisms that form the basis of the maps.In both cases, hypothesis-driven research is clearly key. But perhaps the central message is that if you don’t look at your data first, you might invent solutions that are simply different from how the brain solves a particular problem you care about. When artificially placed on either end of an analytic spectrum, the two approaches expose an epistemological schism: Should we first choose what to look for (computation) or instead interrogate our data (network) before trying to figure out how the brain solves the task?
Fortunately, there are numerous approaches emerging in the literature that afford the discerning scientist the capacity to have their cake and eat it too: That is, the two approaches can be profitably combined to offer better insight into the problem at hand.
For instance, by training artificial networks (such as RNNs) to perform similar switching tasks to those that we scan, we can compare the emergent signatures of the neural network with those seen in the brain, in an effort to determine whether there are commonalities or differences that expose the mechanism of the task. We can also build neuroanatomically plausible models of the brain that can be analyzed like neuroimaging data, though training complex networks to do anything even vaguely like the complex tasks we are interested in can be difficult.
A related strategy is to interrogate neuroimaging data using clustering approaches to reduce the dimensionality of the signal to a more manageable size: At this scale, it’s possible to build toy neural networks to perform the task. Importantly, these toy networks can then be interrogated in manners not possible in a standard human neuroimaging experiment, enabling hypothesis-driven studies to determine the causal mechanisms underlying its function.
These approaches all scratch the surface of a much deeper set of opportunities for modern neuroscience to strike a balance between measurement and hypothesis-driven interrogation of the brain.
Explore more from The Transmitter
A new atlas of abstracts visualizes the field of human brain mapping—where does your work fit?