No two people with autism have the same combination of behaviors, intellectual capabilities or connections in their brains, but these traits can be used to categorize them into four biologically relevant subgroups, according to a new study. Each subgroup correlates with a specific pattern of gene expression.
The subgroups could help researchers identify who is most likely to benefit from experimental therapeutic approaches, such as oxytocin therapy, says study investigator Amanda Buch, a postdoctoral researcher at Weill Cornell Medicine in New York City.
“A lot of the clinical trials for autism have failed,” Buch says. “That’s not that surprising if we think there are different things happening biologically in each of these subtypes.”
Buch and her colleagues took advantage of a wealth of new open-source data, including postmortem gene-expression maps, brain scans and behavioral data from people with and without autism. Although many scientists are leveraging these sophisticated tools to gain new insights into autism, outside researchers say they are impressed by the machine-learning approach Buch and her colleagues used to parse the data in multiple dimensions.
“It’s really amazing that they got four groups,” says Catherine Lord, distinguished professor of psychiatry and education at the University of California, Los Angeles, who helped establish the open-source brain scan database but was not involved in the new study. “There have been a lot of attempts to create subgroups, but it is quite hard.”
Others noted, alongside their praise, that the paper makes bold claims. “When I read it, I said, ‘Wow, these guys cracked autism somehow,’” says Alessandro Gozzi, senior researcher at the Istituto Italiano di Tecnologia in Rovereto, Italy. “Now there will be scrutiny.”
O
ne of the central challenges of understanding, let alone treating, autism is that it is a heterogenous condition. Although people may share some common phenotypic traits, such as repetitive movements or communication difficulties, they each possess their own unique mix of these traits. The condition is associated with more than a thousand genetic variants, most of which have only a small effect.Earlier attempts to partition autism into subgroups based on behavioral traits were largely abandoned after the 2013 publication of the 5th edition of the Diagnostic and Statistical Manual of Mental Disorders (DSM-5). The DSM-5 defined “autism spectrum disorder” as a broad diagnosis and eliminated subtypes, such as Asperger syndrome and the pervasive developmental disorders, that had proven to be unreliable and lacked a biological basis.
“It got so messy, and there was no reliability to the different terms people used,” Lord says. Yet the prospect of using the modern tools of genetics, neuroscience and computing to somehow partition the condition into subgroups remained a long-standing goal for many in the field.
The inspiration for the multidimensional approach initially came from Buch’s doctoral thesis adviser and a collaborator on the study, Conor Liston, associate professor of neuroscience and psychiatry at Weill Cornell Medicine. Liston had previously conducted a brain imaging study to identify depression biomarkers corresponding to four neurophysiological subtypes. Those subtypes, Liston found, could help predict who would best respond to transcranial magnetic stimulation therapy.
“Conor and I had the idea to unravel the heterogeneity of autism and define more biologically associated subtypes,” Buch says.
S
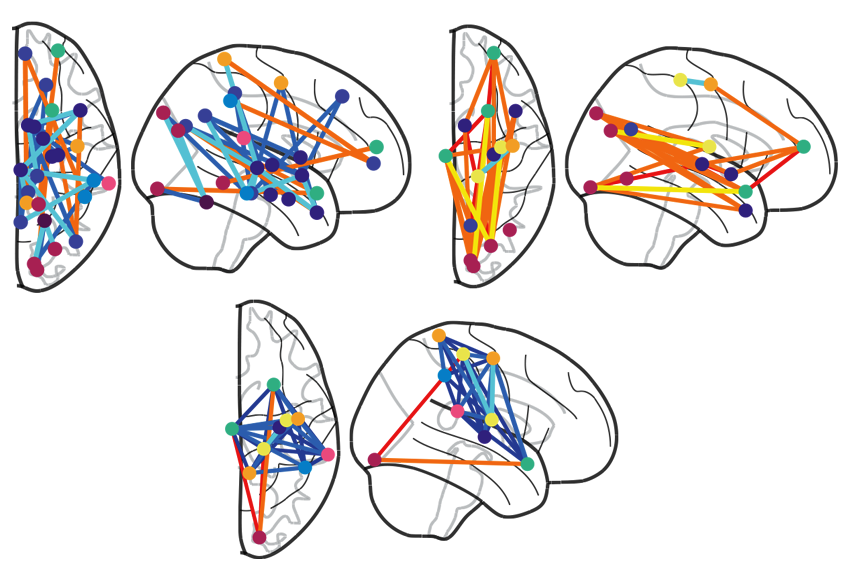
he and her colleagues began by obtaining data from the Autism Brain Imaging Data Exchange, which includes functional MRI data along with verbal intelligence scores and measures of social affect and repetitive behaviors. The researchers then compared resting-state brain activity patterns between 299 people with autism and 907 neurotypical controls to identify atypical patterns of functional connectivity among different brain regions.Their machine-learning technique identified three roughly orthogonal brain-connectivity scores that did the best job of predicting the mix of someone’s verbal intelligence, social affect and repetitive behaviors. The social affect dimension, for instance, ended up being closely linked to connectivity among brain regions associated with social and emotional processing. The verbal intelligence dimension, meanwhile, was associated with connectivity among brain areas related to reading and language.
The study was published on 9 March in Nature Neuroscience.
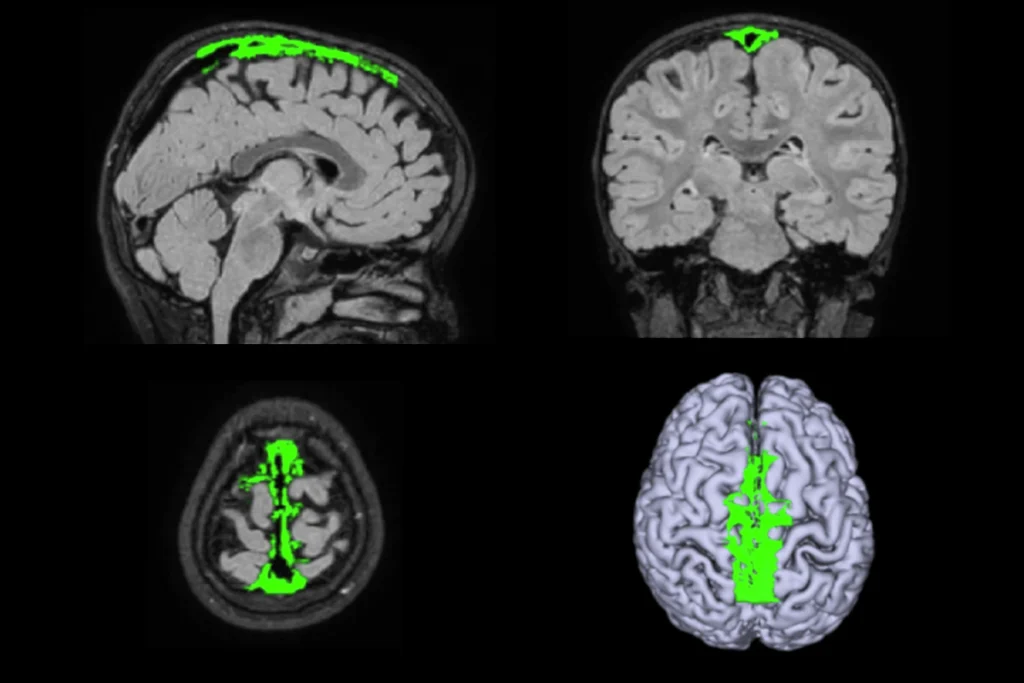
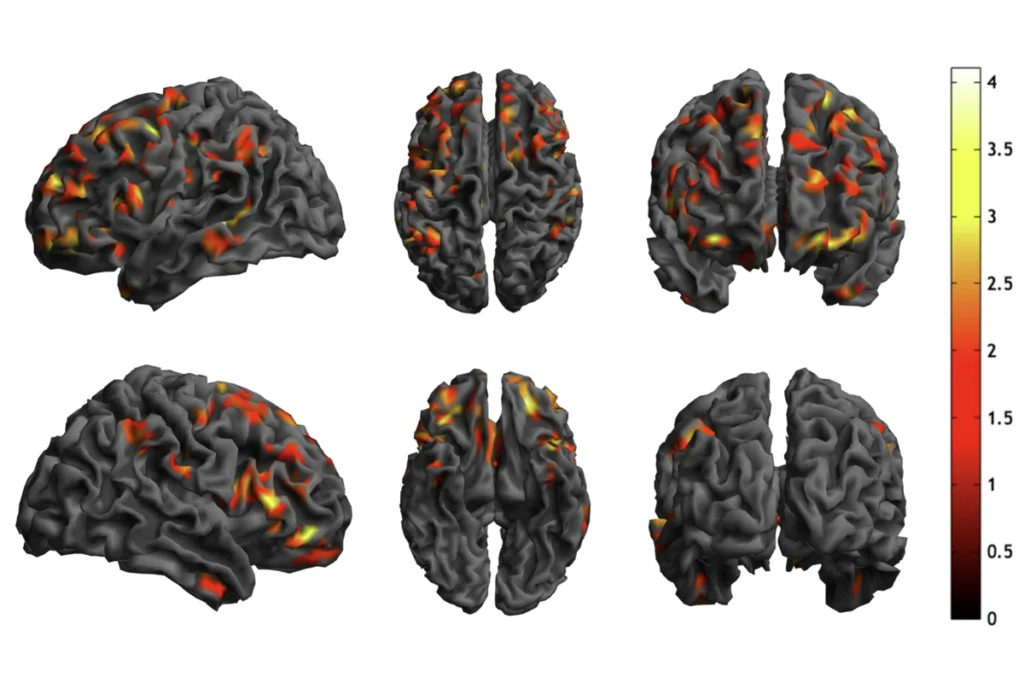
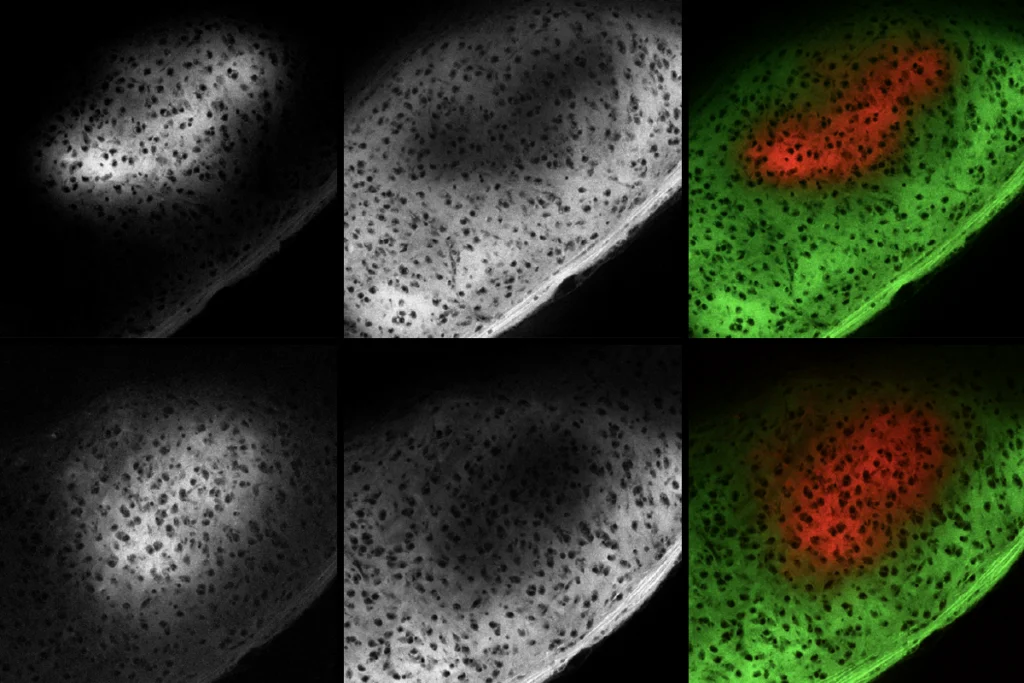
By plotting a person’s score in each of these three brain-behavior dimensions, the researchers were able to identify four distinct clusters, or subgroups. Less than 5 percent of the 4,257 atypical brain connections in autism explained the majority of the differences among these four subgroups. The researchers then used the Allen Human Brain Atlas — which has measured gene-expression patterns from 3,702 brain regions from six neurotypical adults — to identify distinct molecular pathways associated with the atypical activity patterns in each of these subgroups.
In one subgroup, which was characterized by people with high levels of repetitive behaviors, brain activity patterns correlated with decreased expression of the gene HTR1A, which encodes a serotonin receptor and has been previously linked to autism. Another subgroup, which was most challenged in terms of social affect, was associated with the oxytocin receptor. The researchers say they hope to reanalyze data from a failed clinical trial of oxytocin therapy to see if there are indications that it was beneficial for this particular subgroup.
Brain scans might not even be necessary to take advantage of the approach, Buch says. She and her colleagues found that it was possible to assign 226 of the 299 participants in the ABIDE dataset to the correct subgroups using the phenotypic data alone. The ultimate goal, she says, is to identify biomarkers linked to each of these subgroups that would provide even greater accuracy in clinical settings without the use of an MRI machine.
Lord is enthusiastic about that possibility and curious about how stable the subgroup assignment is over a person’s lifetime. “Are they going to be different when they are 60?” she says. “It is relevant to clinical practice.”