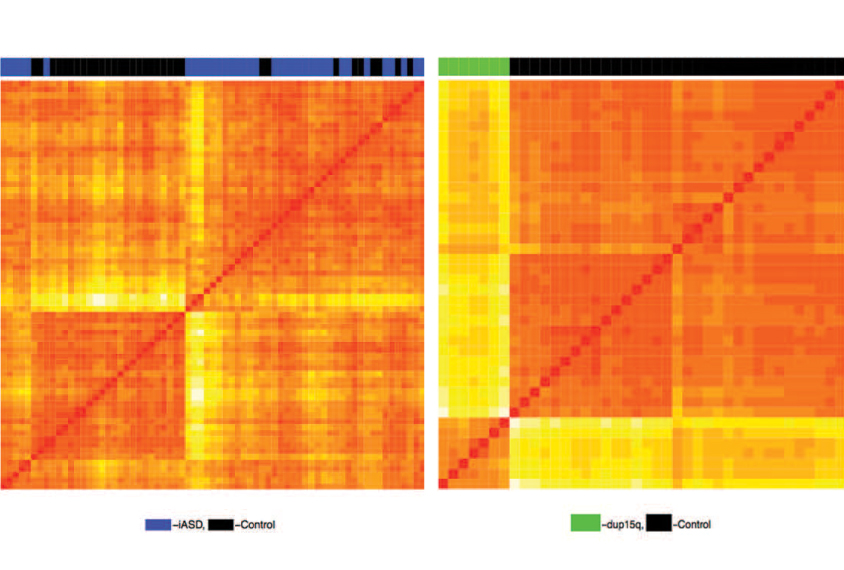
Tag teams: Patterns of chemical tags in brain tissue from people with autism of unknown cause (blue) or from people with a DNA duplication (green) differ from patterns in controls (black).
Different forms of autism may share pattern of chemical tags on DNA
Brain tissues from people with autism of unknown cause and from people with either of two genetic forms of the condition all show similar patterns of methyl groups on DNA.
Brain tissues from people with autism of unknown cause and from people with either of two genetic forms of the condition all show similar patterns of chemical tags on DNA, according to two new studies1,2. The work suggests that a shared biological mechanism underlies different forms of autism.
The studies provide the most thorough look to date at the distribution of chemical tags called methyl groups, which tend to turn off gene expression, in autistic people. The first study looked at DNA methylation in the largest-yet sample of autism brains; the second is the first to investigate methylation across the genome in brain tissue from autistic people.
The samples came from people with either idiopathic autism (autism with no known cause); a genetic condition called dup15q syndrome, in which a region of chromosome 15 is duplicated; or Rett syndrome, a condition related to autism.
“You actually see quite a consistent effect,” says Jonathan Mill, professor of epigenetics at the University of Exeter in the United Kingdom, who led the large study. “It suggests at a molecular level there is some kind of convergent pathology.”
Both studies identify similar patterns of methylation, although certain distinctions are also evident. The commonalities congregate in or near genes that operate at neuronal junctions, or synapses, and in genes that regulate the immune system.
“These analyses allow us to ‘zoom out’ and look at DNA methylation marks genome-wide in a single system,” says Rosanna Weksberg, professor of molecular and medical genetics at the University of Toronto in Canada, who was not involved in the research.
The new reports align with smaller studies of DNA methylation in autism, which also show broad similarities among different forms of autism.
“In my view, they bring a closure to this avenue of research,” says Evan Elliott, assistant professor of molecular and behavioral neuroscience at Bar-Ilan university in Israel, who was not involved in either study. “Basically, everyone is finding something which is pretty similar.”
Identifying tags:
Mill’s team analyzed methylation patterns in postmortem brain tissue from 36 people with idiopathic autism, 7 with dup15q syndrome and 38 controls. They looked at three brain regions — the prefrontal cortex, temporal cortex and cerebellum.
The prefrontal and temporal cortices of people with idiopathic autism show different methylation patterns from those of controls. These brain regions also show distinctive patterns of methylation in dup15q brain tissue, mostly in the genomic areas surrounding the mutation, in line with previous work3. (The researchers did not find any autism-specific patterns in the cerebellum.)
Researchers often assume that epigenetic changes such as DNA methylation occur independently of mutations, says Mill, but the new work suggests that mutations can strongly influence the patterns.
Still, the methylation patterns in the dup15q and idiopathic autism brains overlap significantly in the cortex — predominantly in synapse and immune-system genes. Researchers reported the results 1 July in Human Molecular Genetics.
The study is part of a project to characterize various processes that regulate gene expression, using the same tissue samples. Tissue from people with idiopathic autism and those with dup15q syndrome also show similar patterns of gene expression, as well as in acetylation of histones, another kind of chemical modification4. The researchers are aligning the maps to see whether the sets of differences occur in similar places in the genome.
What is needed now is to determine how these patterns affect brain function, Elliott says.
Same story:
One limitation of the study is that the arrays Mill’s team used to assess methylation provide information about just 2 percent of the genome. They may not capture the areas where most variations in methylation patterns are likely to lie, says Janine LaSalle, professor of medical microbiology and immunology at the University of California, Davis, who led the second study.
La Salle’s team used a more comprehensive technique called bisulfite sequencing to assess methylation patterns in frontal-cortex tissue from 17 autistic people and 6 people with Rett syndrome, and from the visual cortex of 5 people with dup15q syndrome. They compared this tissue with that from 21 controls.
They mapped methylation in fragments about 700 nucleotides long to identify subsections in which methylation patterns differ between autistic people and controls. They overlaid these sections onto publicly available maps of gene expression in various cell types. The overlap revealed which parts of genes in the sections are methylated and the cell type each section comes from.
The sections of the genome with altered methylation are not the same in autism, Rett and dup15q, the researchers found. But the alterations converge on similar genes and pathways in all three conditions. The work appeared in June in Cerebral Cortex.
The differences frequently occur in or near genes that play a role in nervous-system development and are especially enriched in microglia, the brain’s immune cells, which prune synapses. The results broadly align with those from the other study, LaSalle says.
“If you use different approaches and you end up telling the same big-picture story, I think that [adds weight to] both papers,” she says.
All postmortem brain studies are limited by a lack of high-quality tissue, so Mill says he would like to pool his results with LaSalle’s for a more robust sample. “It would be really interesting to put these two datasets together and really see what commonalities and overlaps in signal there are,” he says.
References:
Explore more from The Transmitter

Still no proof for facilitated spelling methods

