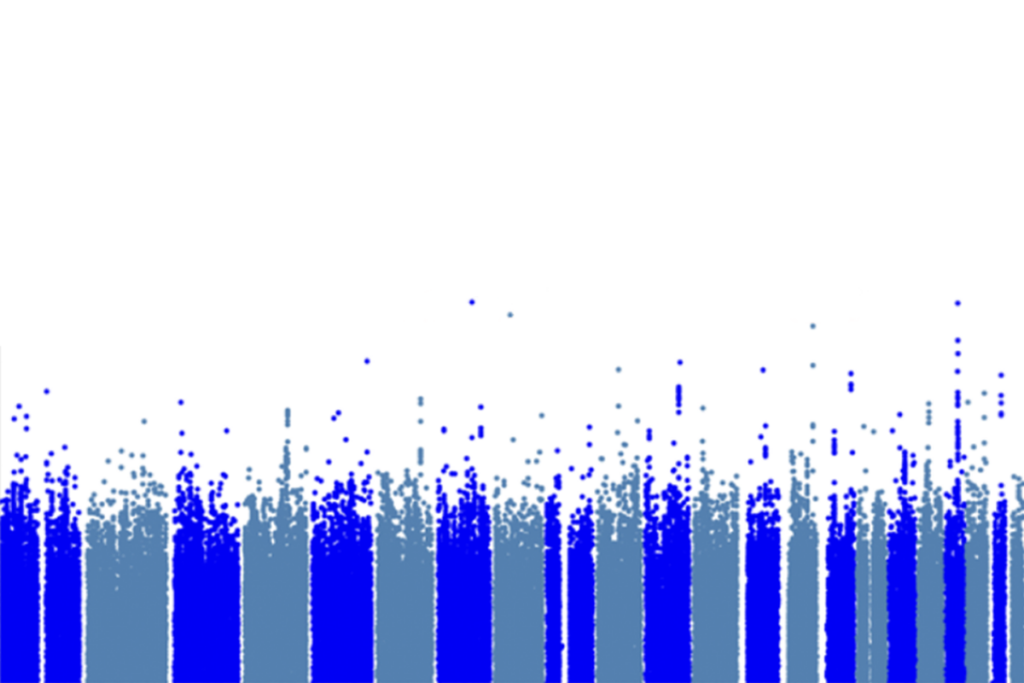
Shared genetic risk
A large study identifying common genetic variations that contribute to five different psychiatric disorders, including autism, sheds light on the disorder’s genetic architecture, says Benjamin Neale.
Common genetic variations can raise risk for five different psychiatric disorders: attention deficit hyperactivity disorder (ADHD), autism, bipolar disorder, major depressive disorder (MDD) and schizophrenia.
On 21 February in The Lancet, the Cross-Disorder Group of the Psychiatric Genomics Consortium (PGC), of which I am a member, reported on a combined genome-wide association study (GWAS) across the five different conditions1.
For autism, this work represents promising new leads for understanding the genetics of the disorder.
Our team analyzed 33,332 individuals with one of the disorders and 27,888 controls using two primary analytic strategies: looking at disease risk linked to individual variants, as well as sets of variants overall. We also looked at whether variants are linked to individual disorders or to a combination.
Individual variants are helpful to determine whether there is shared biology across different disorders. The composite of variants allows us to assess the strength of the genetic overlap among the disorders.
The study identified four regions housing variants that are significantly linked to the various disorders. In three of these four regions, the data suggest that the effect is shared for all five diseases. In the fourth region, a variant in the calcium channel gene CACNA1C appears specific to bipolar disorder and schizophrenia.
Common links:
A number of GWAS have identified variants linked to bipolar disorder and schizophrenia2,3 but not to autism. In the new study, we analyzed ten different schizophrenia-linked variants; four of these are specific to schizophrenia, and the other six are more diverse. Strikingly, three of these six seem to be linked exclusively to schizophrenia and autism, suggesting that some of the genetic effects that confer risk for schizophrenia also do so for autism.
The findings aren’t a surprise — a number of studies have linked the two disorders. Autism was once considered a childhood form of schizophrenia, and a family history of schizophrenia raises the risk of autism. Certain genetic variants are linked to both disorders, including deletion of 16p11.2. (Some research refutes a strong genetic link.)
However, most studies have focused on copy number variations, deletions or duplications of chunks of DNA that are typically rare. The new study provides the first evidence of shared genetic risk across the genome for common, single-letter variants.
Overall, the genetic link between autism and the other disorders is less dramatic than it is for, say, bipolar disorder and schizophrenia, which show extensive genetic overlap. But the evidence suggests further sharing between autism and schizophrenia beyond the handful of loci described above.
The findings strengthen the evidence that genetic risk factors occur across all levels of the genetic architecture, including both common and rare variants, and suggest that efforts to understand schizophrenia may yield insights into autism — and vice versa.
What’s more, the new results provide further evidence that common variants play a role in autism, contributing up to 40 percent of the genetic risk of autism.
This work also underscores the importance of large sample sizes for identifying genetic variants specific to autism. That in turn will drive forward the next round of biological studies.
Benjamin Neale is assistant professor in the Analytic and Translational Genetics Unit at Massachusetts General Hospital in Boston.
References:
1: Cross-Disorder Group of the Psychiatric Genomics Consortium. Lancet 381, 1371-1379 (2013) PubMed
2: Ripke S. et al. Nat. Genet. 43, 969-976 (2011) PubMed
3: Sklar P. et al. Nat. Genet. 43, 977-983 (2011) PubMed
Explore more from The Transmitter

Remembering Annette Dolphin, who helped explain gabapentin’s effects