Gene fix: In-vivo base editing can mitigate autism-like effects in mice carrying a variant of the CHD3 gene, according to a new paper. The variant, found in people with Snijders Blok-Campeau syndrome, causes altered sociability, cognition and motor coordination in mice. A TadA-embedded adenine base editor, delivered brain-wide via an adeno-associated virus, restored CHD3 protein levels and ameliorated the behavioral effects. The findings, the researchers write, “demonstrate that precise single-base correction in the postnatal brain can restore protein dosage and function, thereby offering a framework for the treatment of monogenic neurodevelopmental disorders.”
In-vivo base editing in a mouse model of autism, and more
Here is a roundup of autism-related news and research spotted around the web for the week of 23 February.
By
Jill Adams
24 February 2026 | 2 min read
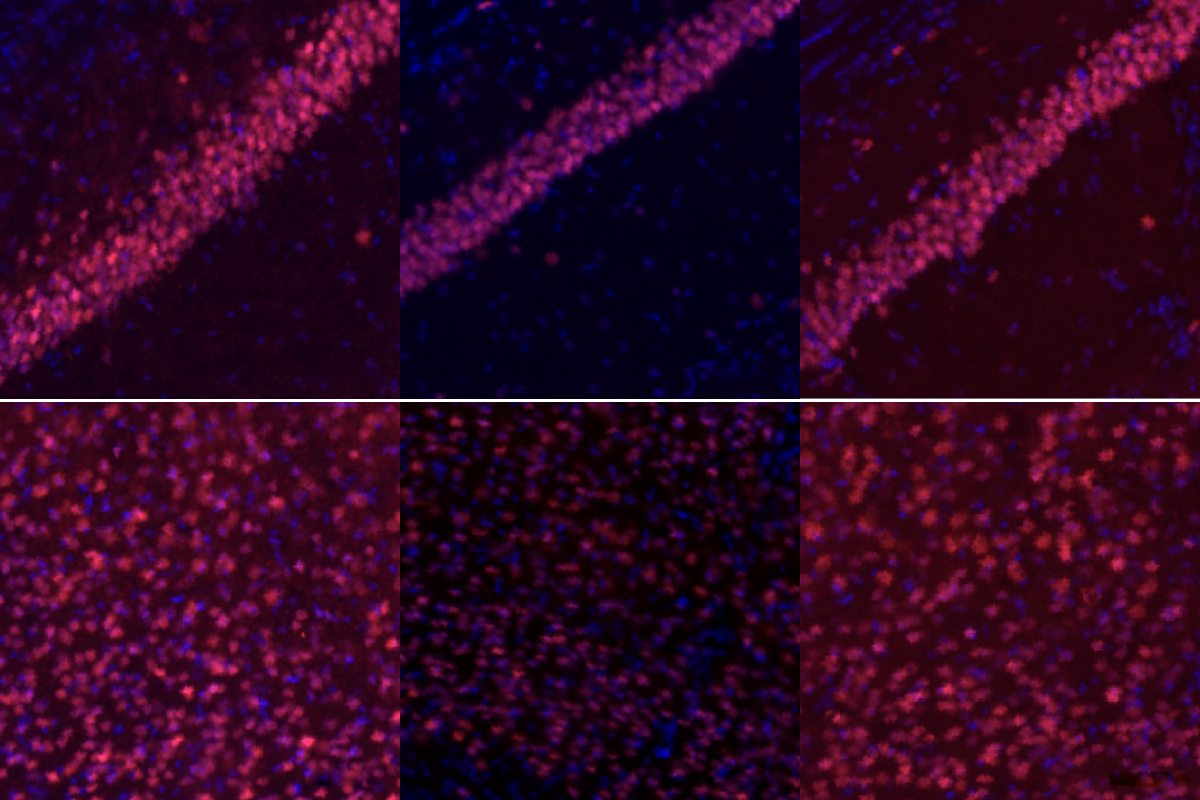
Replacement parts: Mice carrying a CHD3 variant have lower CHD3 protein levels (middle panels) than wildtype mice (left panels) in two brain regions (hippocampus, top row; somatosensory cortex, bottom). In-vivo base editing restores the protein (right panels).
Autism research spotted this week:
- “Characterizing features of the genetic architecture underlying autism from a multi-ancestry perspective” medRxiv
- “Up-regulation of Minibrain/DYRK1A contributes to macrocephaly and brain overgrowth in a Drosophila model of fragile X syndrome” Proceedings of the National Academy of Sciences
- “Genome-wide significance reconsidered: Low-frequency variants and regulatory networks in autism” medRxiv
- “Deficiency of the histone lysine demethylase KDM5B causes autism-like phenotypes via increased NMDAR signalling” bioRxiv
tags:
Explore more from The Transmitter
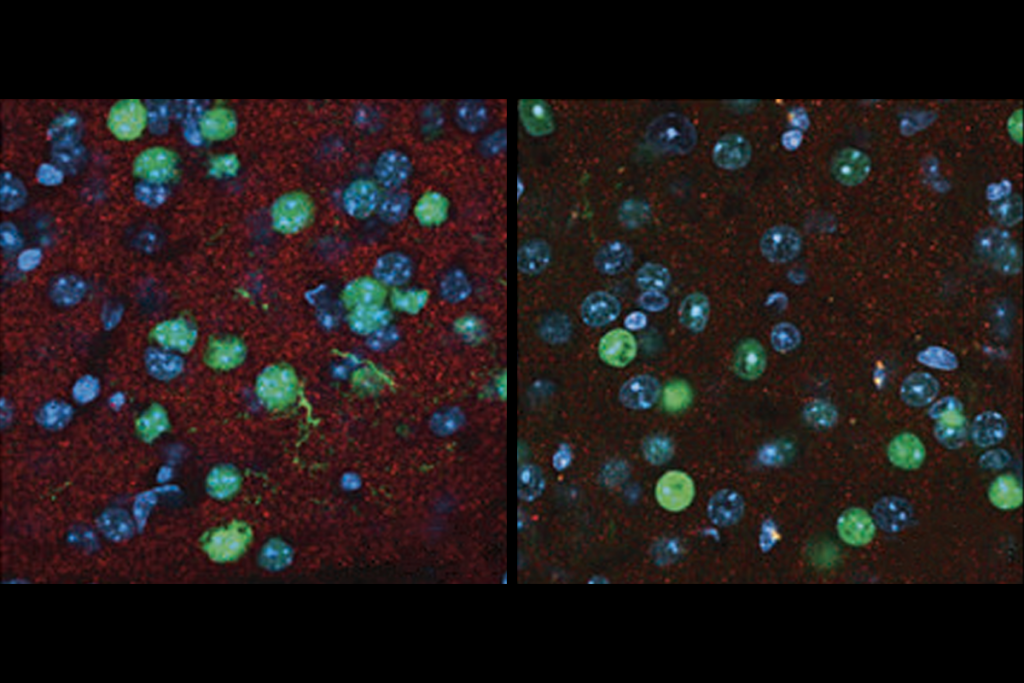
SHANK3 deficiency and behavior in mice; and more
By
Jill Adams
25 November 2025 | 2 min read
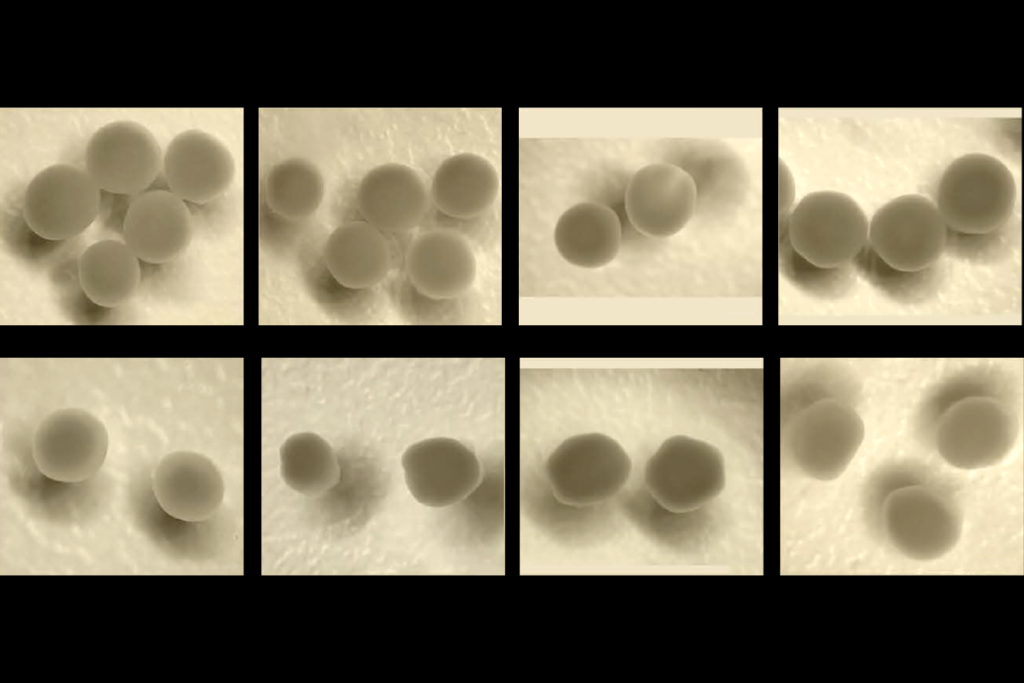

Parsing phenotypes in people with shared autism-linked variants; and more
By
Jill Adams
14 October 2025 | 2 min read
New method identifies two-hit genetic variation in autism; and more
By
Jill Adams
28 October 2025 | 2 min read
Cite this article: