New mouse model mimics many symptoms of autism
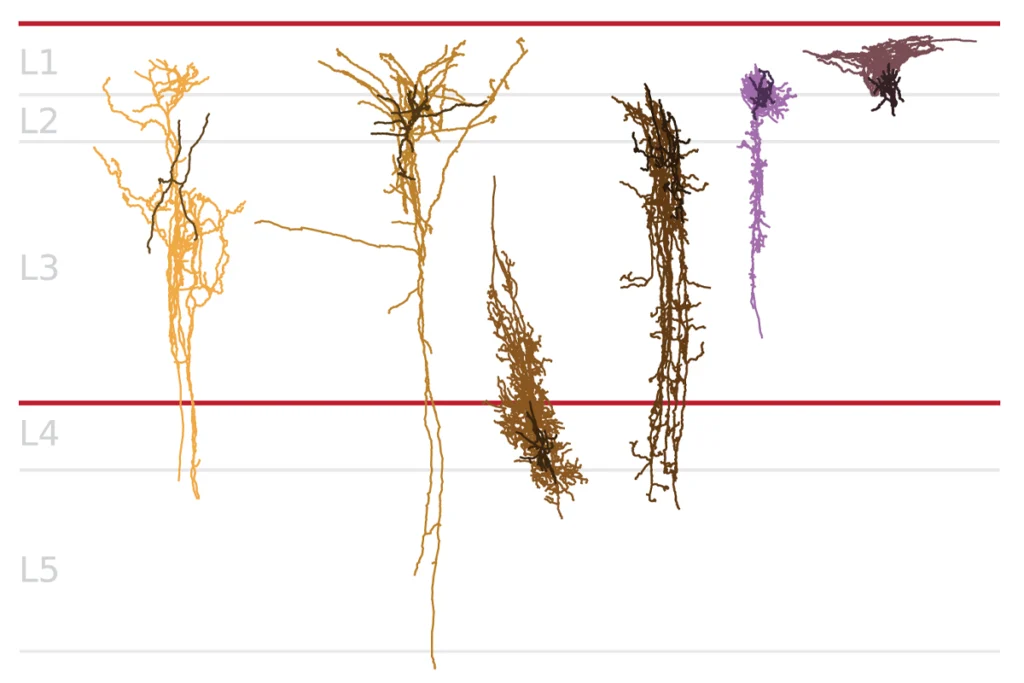
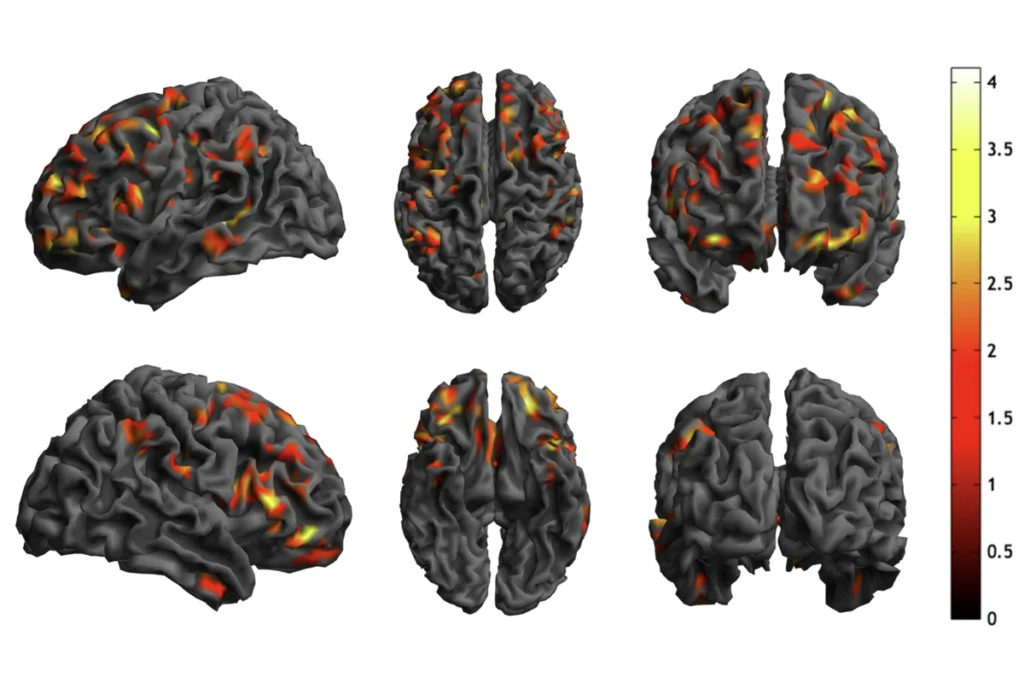
Mice lacking the autism-linked gene CNTNAP2 show many of the behaviors associated with the disorder, and exhibit brain circuit disruptions similar to those seen in people who carry mutations in the gene.
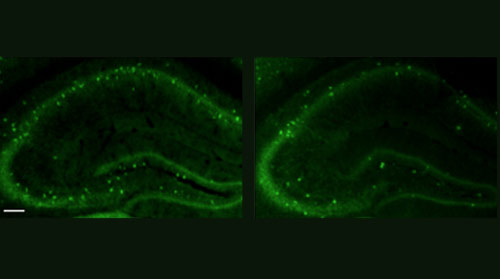
Mice lacking the autism-related gene CNTNAP2 also show brain circuit disruptions similar to those seen in people who carry mutations in the gene, the study found.
What’s more, risperidone, a drug often used to treat the symptoms of autism, ameliorates some behaviors, including hyperactivity and repetitive behavior, in these mice.
The researchers presented an early look at the behavioral findings at the 2011 International Meeting for Autism Research.
The study is clearly a landmark in autism research and cause for optimism, says Nicholas Gaiano, associate professor of neurology at Johns Hopkins University School of Medicine in Baltimore, who was not involved with the work.
“Not only does the work further cement a role for CNTNAP2 in autism biology from a behavioral standpoint, but it provides a beautiful example of how disruption of an [autism]-associated gene can profoundly disrupt brain development,” Gaiano says.
Mutations in CNTNAP2 are associated with a higher risk of autism2,3. People who have inheritedmutations in both copies of the gene have a rare disorder called cortical dysplasia-focal epilepsy syndrome, or CDFE. The syndrome is characterized by epileptic seizures, loss of language, intellectual disability and, in some 70 percent of cases, autism.
Some genetic variants of CNTNAP2 are also associated with specific language impairment, a disorder not accompanied by intellectual disability, autism or physical problems such as hearing loss.
The faithfulness with which the new model recapitulates many of the features of autism came as a surprise to the investigators.
“I thought it would be a great system for studying how this gene affects neural development,” says lead investigator Daniel Geschwind, chair of human genetics at the University of California, Los Angeles School of Medicine.
The mice offer that and much more: They are hyperactive, have social and vocal communication deficits and repetitive behavior, all hallmarks of autism.
“I’ve been somewhat conservative about making parallels between human behavior and mouse behavior,” says Geschwind. “I’m much more optimistic about using mice after seeing this.”
Signaling imbalance:
References:
Recommended reading
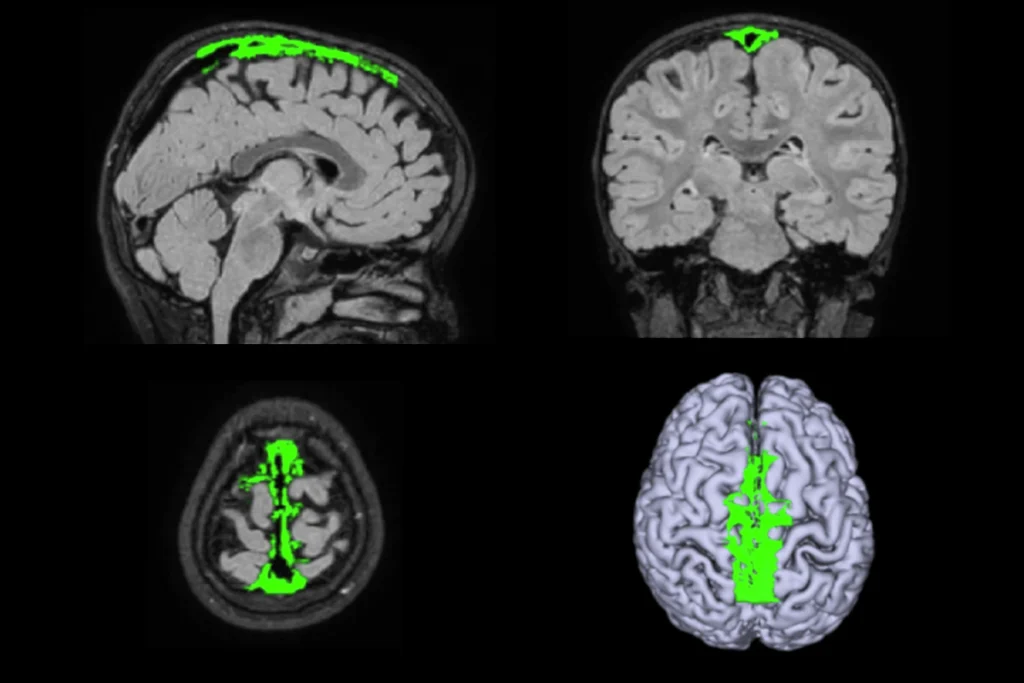
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
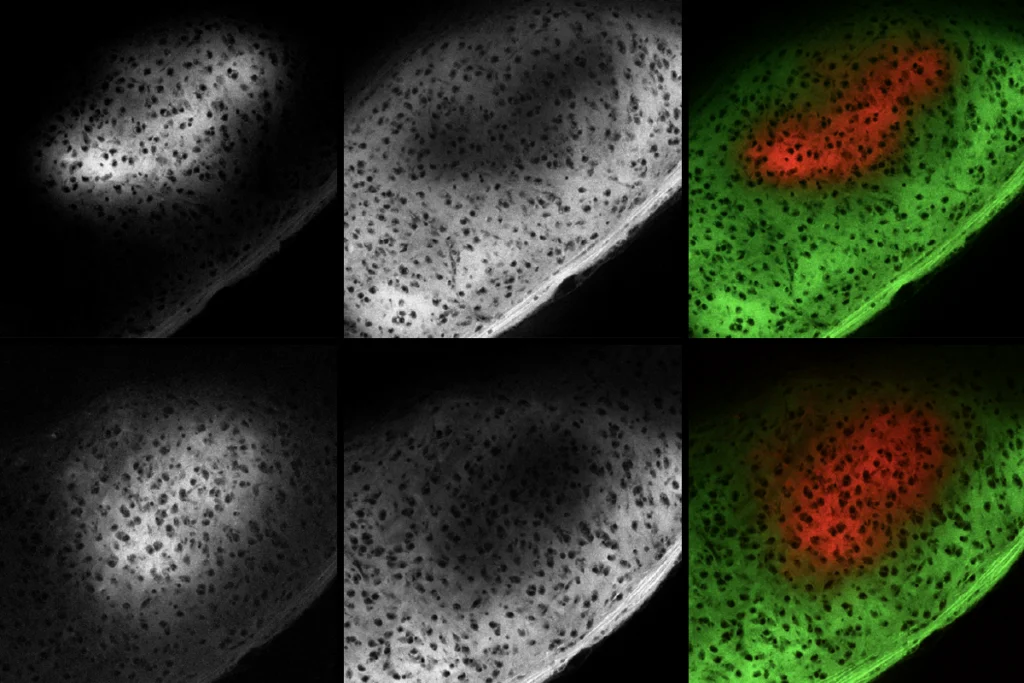
Microglia’s pruning function called into question