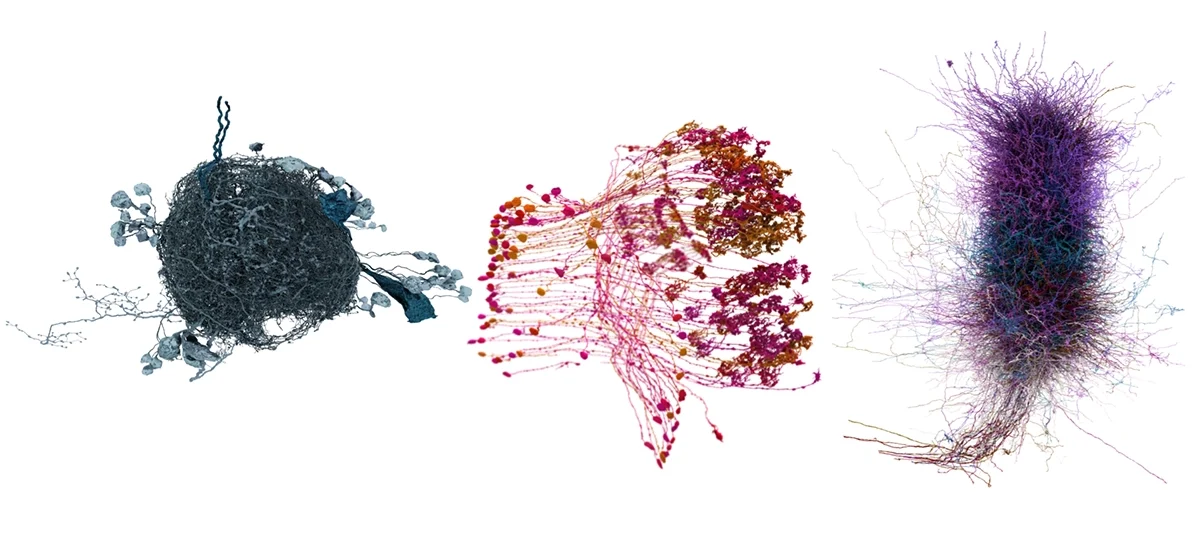
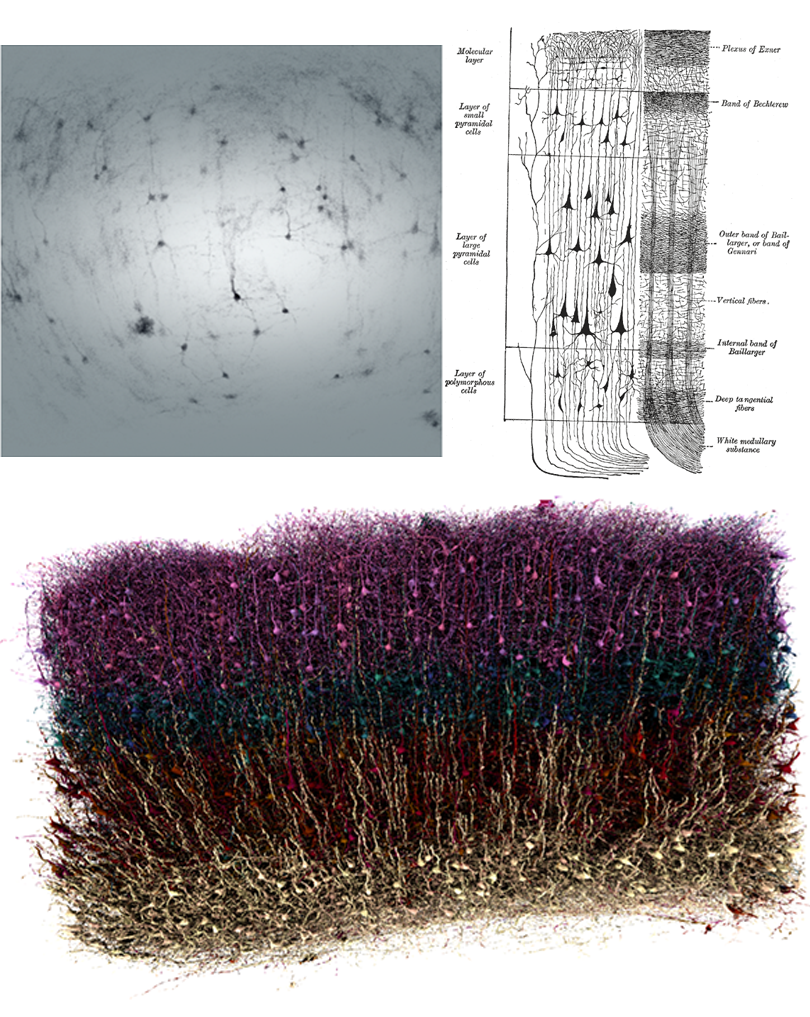
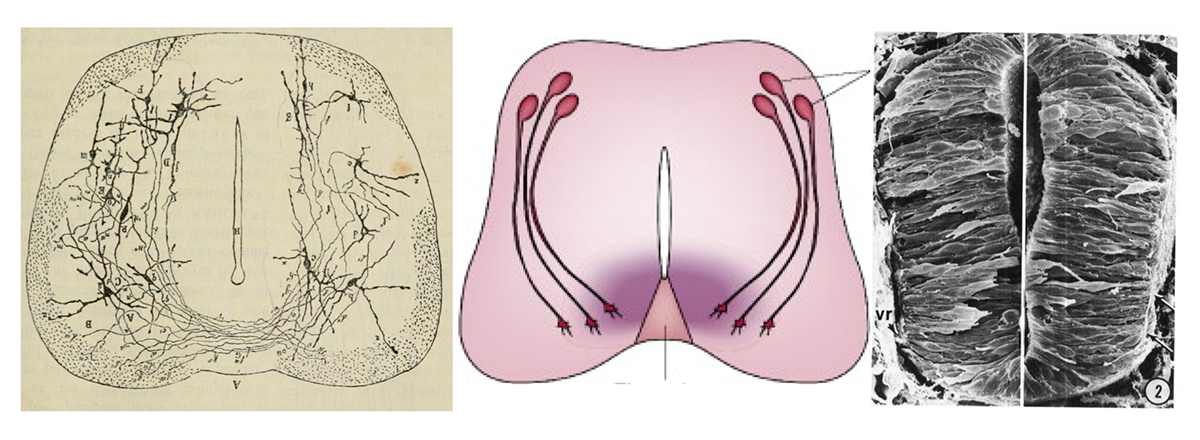
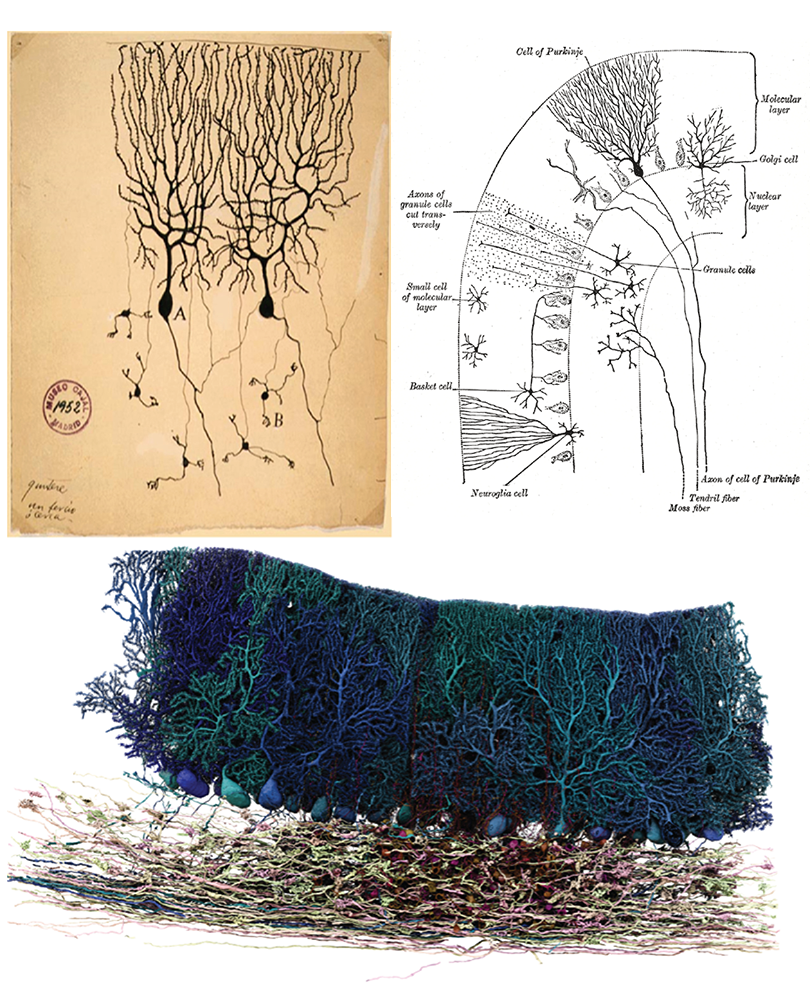
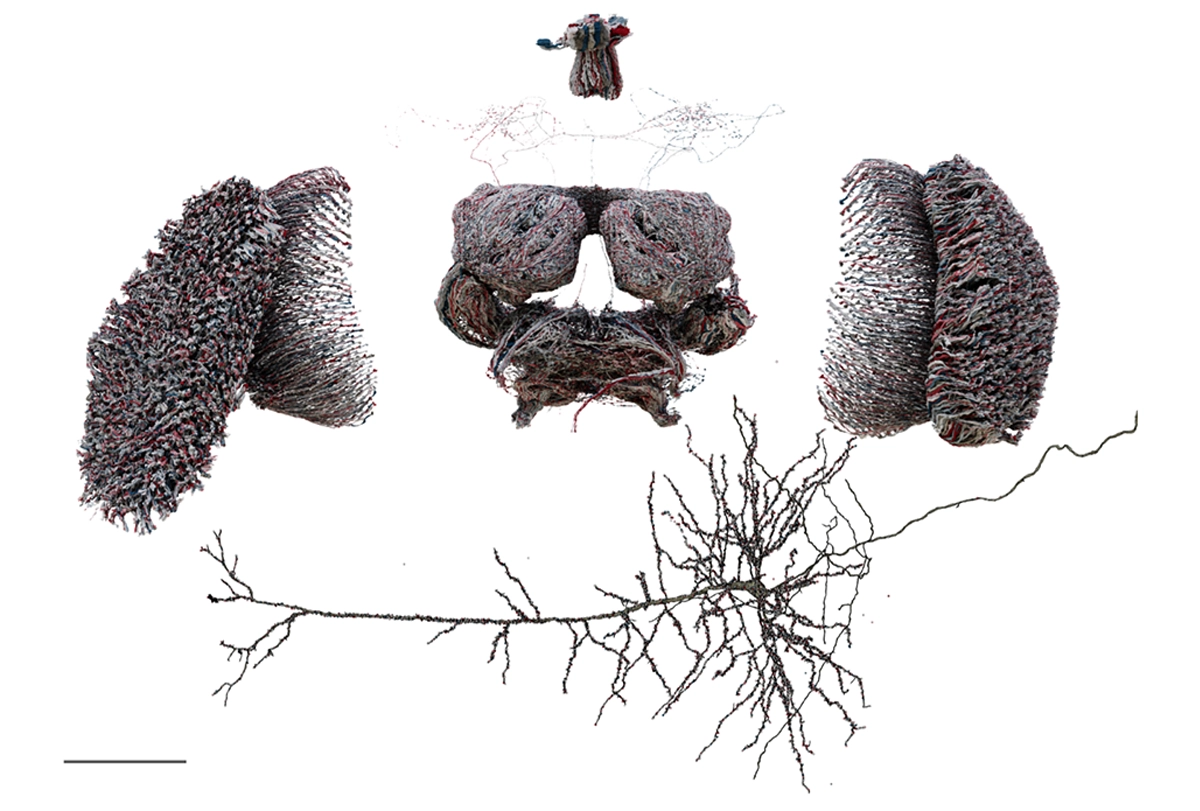
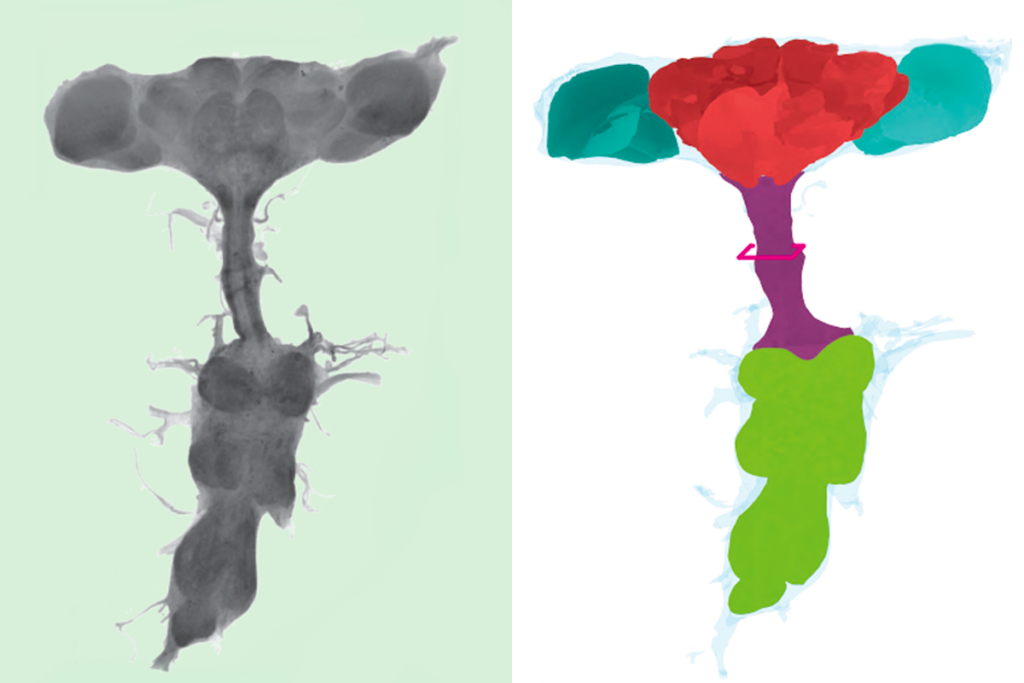
If I ask you to picture a group of “neurons firing,” what comes to mind? For most people, it’s a few isolated neurons flashing in synchrony. This type of minimalist representation of neurons is common within neuroscience, inspired in part by Santiago Ramón y Cajal’s elegant depictions of the nervous system. His work left a deep mark on our intuitions, but the method he used—Golgi staining—highlights just 1 to 5 percent of neurons.
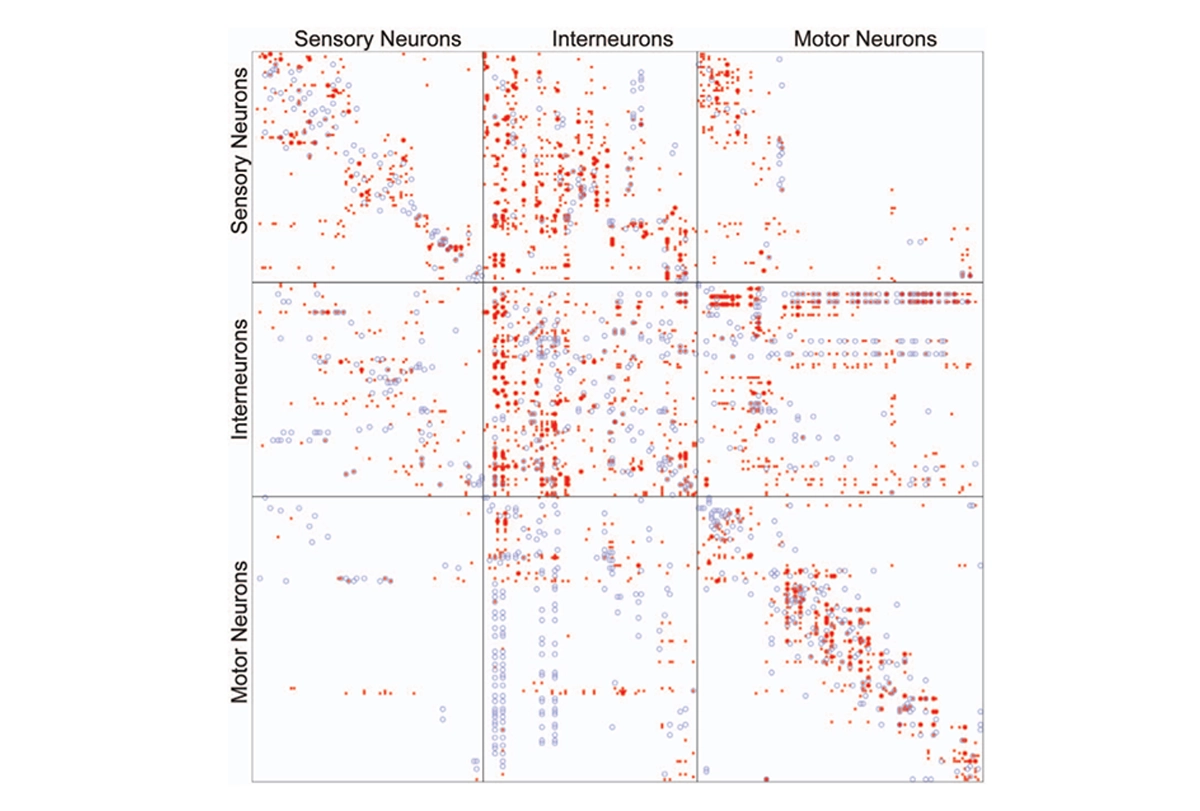
More than a century later, researchers have mapped out brain connectivity in such detail that it easily becomes overwhelming; I vividly recall an undergraduate neurophysiology lecture in which the professor showed a wiring diagram of the primary visual cortex to make the point that it was too complex to understand.