Exploring the hidden world of inhibitory brain signals
Scientists have discovered more than 100 new proteins at junctions in the brain that dampen neuronal activity.

Billions of years ago, a Mars-sized body slammed into Earth, dislodging a spinning ring of debris that formed our moon. As the moon’s rotation slowed, it became phase-locked relative to the Earth, keeping its far side hidden from our view. In October 1959, the Soviet spacecraft Luna 3 took the first picture of the far side of the moon. In the same month, Edward George Gray, a pioneer in studying brain ultrastructure, took some of the first electron microscopy images of connections between brain cells.
Gray’s pictures revealed two types of junctions, or synapses, between neurons. The symmetrically shaped synapses are ‘inhibitory,’ meaning they tend to depress activity in neurons. The synapses that appear asymmetric are ‘excitatory;’ they boost brain activity.
Since this discovery, researchers have extensively probed the latter type of synapses. These junctions sport about 1,000 proteins involved in functions ranging from remodeling the synapse to trafficking proteins across it. This basic molecular framework has served as a platform to study how excitatory synapses develop and function.
We have also learned that mutations, and DNA duplications and deletions associated with autism, affect the way many of these proteins work. By contrast, inhibitory synapses are difficult to isolate biochemically, and so relatively little is known about them.
In many respects, the molecular composition of inhibitory synapses is like ‘the far side of the moon.’ The proteins remain largely mysterious and uncharted. But last year, researchers made major inroads into this molecular territory. They found strong hints that mutations in some of these proteins contribute to autism.
Brakes in the brain:
Autism is extremely heterogeneous, genetically and behaviorally. As a result, constructing a unifying theory of the condition remains difficult. One intriguing idea is that autism arises from too much excitation relative to inhibition of neuronal activity.
Early support for a so-called signaling imbalance in autism comes from the observation that many people with autism also have epilepsy. Recordings of electrical activity during sleep in the brains of children with autism also suggest that 50 to 70 percent of these children have hyperexcitable neural networks1.
Postmortem studies of people with autism suggest these effects are due to decreased signaling of cells via the inhibitory molecule gamma-aminobutyric acid (GABA). For example, the cerebral cortex (outer portion of the brain) in those with autism contains low levels of enzymes that synthesize GABA2. Some evidence suggests that levels of receptors for GABA are also unusually low in the brains of people with autism3.
Consistent with these studies, mutations linked to autism can abolish inhibitory signals in mice. Tuberous sclerosis, a syndromic form of autism, arises from dysfunction of the TSC1 gene. Mice lacking the gene show a deficit in signaling at inhibitory synapses in the hippocampus and are susceptible to seizures.
The autism-related condition fragile X syndrome is also associated with abnormalities in inhibitory signaling. Last year, researchers found that disruption of the gene PX-RICS, which facilitates the trafficking of a GABA receptor at inhibitory synapses, is linked to Jacobson syndrome, another condition that overlaps with autism. Loss of this gene in mice results in a decrease in GABA signaling and behaviors reminiscent of autism4.
Protein puzzle:
These data all indicate that problems with GABA signaling underlie autism. Yet our lack of understanding of the proteins (and their associated genes) at the inhibitory synapse has hampered progress toward pinpointing the pathways that lead to these problems.
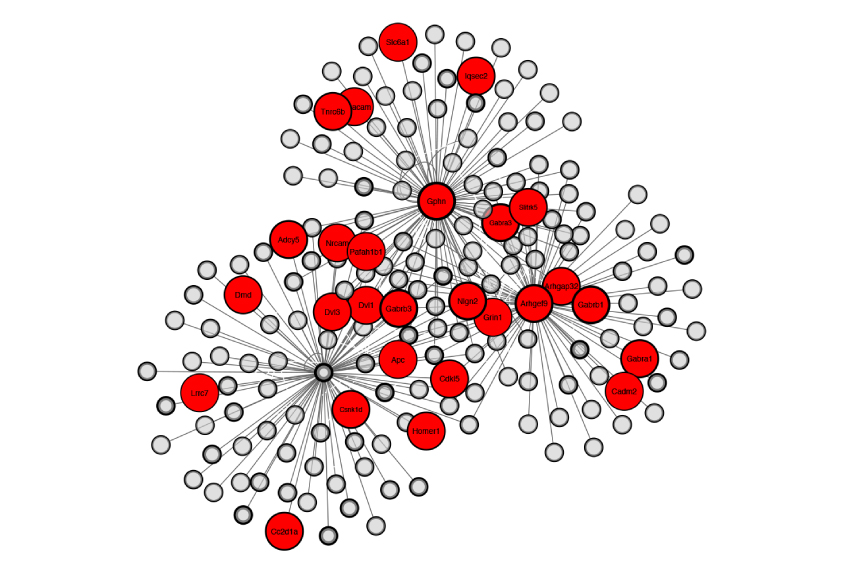
Two teams of researchers addressed this critical question last year. Both outlined molecular approaches for tagging and purifying the proteins at different locations within inhibitory synapses5,6. Using these approaches, they identified a number of proteins that govern the development and function of the synapses.
This new ‘inhibitory synaptic proteome’ — essentially, a library of the proteins at this type of junction — covers virtually all of the proteins we knew about, along with more than 100 we did not. The known proteins include GABA receptors and proteins that influence those receptors such as PX-RICS.
Among the new proteins are those that shore up the physical structure of the synapse, act as enzymes, or span its outer membrane to presumably connect or communicate with other neurons. Many are of unknown function.
Importantly, approximately 15 percent of the proteins at the inhibitory synapse are implicated in autism.
Identifying the proteins at inhibitory synapses is a breakthrough in basic neuroscience —not unlike the breakthrough in space exploration that came from Luna 3’s snapshots. With these new proteomic maps in hand, we can explore the molecular landscape of the inhibitory synapse.
Scott Soderling is associate professor of cell biology and neurobiology at Duke University in Durham, North Carolina. Akiyoshi Uezu is a senior research associate in his lab.
References:
- Lewine J.D. et al. Pediatrics 104, 405-418 (1999) PubMed
- Fatemi S.H. et al. Biol. Psychiatry 52, 805-810 (2002) PubMed
- Fatemi S.H. et al. J. Autism Dev. Disord. 39, 223-230 (2009) PubMed
- Nakamura T. et al. Nat. Commun. 7, 10861 (2016) PubMed
- Loh K.H. et al. Cell 166, 1295-1307 (2016) PubMed
- Uezu A. et al. Science 353, 1123-1129 (2016) PubMed
Explore more from The Transmitter

Remembering Annette Dolphin, who helped explain gabapentin’s effects