When Maxwell Elliott’s latest research paper began making the rounds on Twitter last June, he wasn’t sure how he felt.
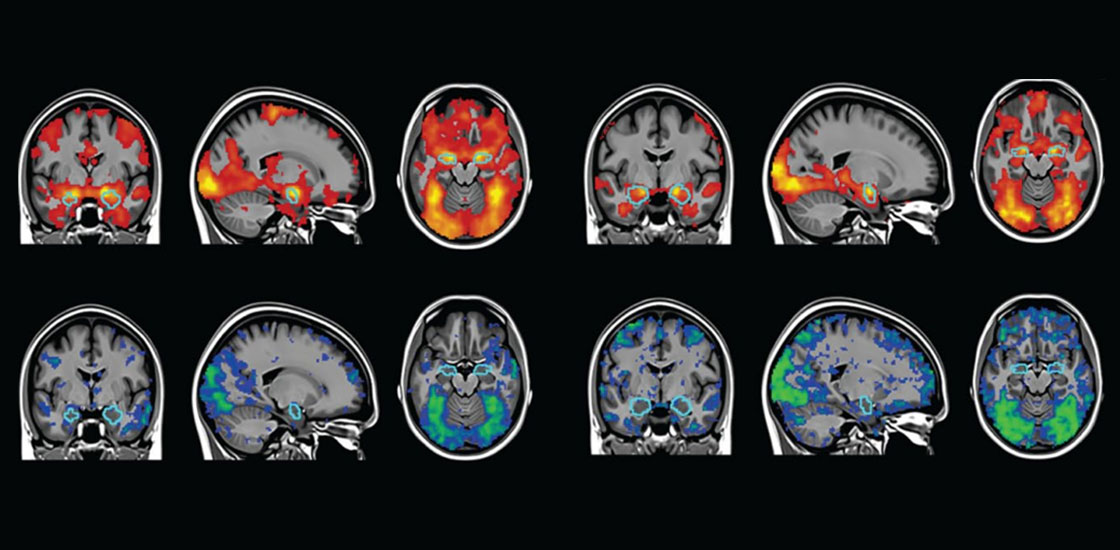
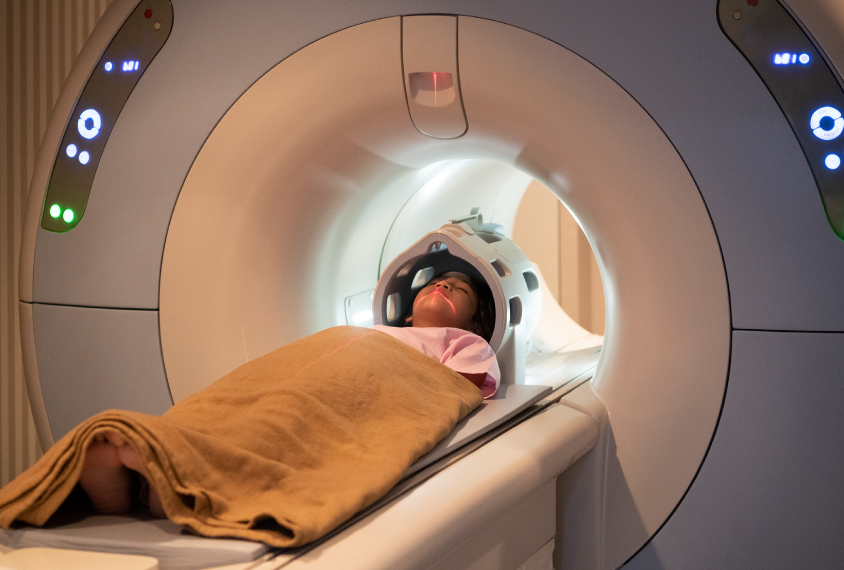
Elliott, a graduate student in clinical psychology in Ahmad Hariri’s lab at Duke University in Durham, North Carolina, studies functional magnetic resonance imaging (fMRI) and how it can be used to better understand neurological conditions such as dementia and autism.
He was excited that this “nitty-gritty” aspect of the field, as he describes it, was garnering a bit more attention, but the reason for the buzz disappointed him: A news outlet had picked up the story and run it with an overstated headline: “Duke University researchers say every brain activity study you’ve ever read is wrong.”
“They seemed to just totally misunderstand it,” Elliott says.
His study, published in Psychological Science, did not discount 30 years of fMRI research, as the headline suggested. It did, however, call into question the reliability of experiments that use fMRI to tease out differences in how people’s brains respond to a stimulus. For certain activities, such as emotion-processing tasks, individual differences in brain activity patterns do not hold up when people are scanned multiple times, months apart, Elliott and his colleagues had shown.
Elliott had been worried about the response from researchers who use such patterns to try to distinguish between people with and without certain neuropsychiatric conditions, including autism. The sensationalized reporting only deepened that fear.
Many in the brain imaging field jumped to defend Elliott and his colleagues and condemned the news outlet’s mischaracterization, but others took issue with the study and what they saw as an overgeneralization of the limits of fMRI. On both sides, the idea that fMRI has a reliability problem struck a chord — one that reverberated far and wide in the Twittersphere.
For all the chatter last year, concerns about brain imaging research are nothing new. Critics have previously claimed, misguidedly, that fMRI is so useless that it can find signal in a dead salmon and, more reasonably, that even breathing can distort brain scan results beyond usability.
Regardless, many brain imaging researchers are optimistic about what the tools can do. They don’t dismiss the problems other scientists have raised, but they tend to view them as growing pains in what is still a relatively new field. Some problems, particularly those that arise through data collection and analysis, can likely be solved; some may be more endemic to the methods themselves. But none are as dire as many tweets and headlines proclaim.
“It doesn’t hurt to remind people that you’ve got to keep those limitations in mind, no matter how advanced the field gets,” says Kevin Pelphrey, professor of neurology at the University of Virginia Brain Institute in Charlottesville.
Flushing the pipeline:
Some problems with imaging studies start right with data collection. The Autism Brain Imaging Data Exchange (ABIDE) dataset, for example, launched with a relatively small number of scans — from 539 autistic people and 573 non-autistic people — at 17 different sites, and an early analysis of the dataset found wide variability based on where the scans took place. A 2015 study showed that this variability could even lead to false results.
Since then, the researchers who run ABIDE have added another 1,000 brain scans and use a total of 19 sites. The larger sample sizes should improve some of the replicability issues across the scans, the team has said. They also plan to meet as a group to discuss how to standardize data collection.
Other problems arise when imaging data reach the analysis stage. In May 2020, just weeks before Elliott and his colleagues published their study, a team from Tel Aviv University in Israel published a paper in Nature showing how different pipelines for analyzing fMRI data can also lead to widely variable results.
In that work, 70 separate research teams analyzed the same raw fMRI dataset, each team testing the same nine hypotheses, but because no two teams used the exact same workflow for analyzing the scans, they all ended up reporting different ‘findings.’
Workflow issues also plague structural MRI scans, which reveal brain anatomy instead of activity. Although this form of MRI is more reliable than fMRI from scan to scan, according to the study by Elliott and his colleagues, one unpublished investigation showed how 42 groups trying to trace white-matter tracts through the same brain rarely produced the same results. (A 2017 study found that groups could successfully identify 90 percent of the underlying tracts, but not without producing a large number of false positives.)
Workflow inconsistencies are a major reason brain imaging results have been challenging to reproduce, Pelphrey says. Each decision made during the analysis of a set of images — such as where to set a threshold for what constitutes brain activity, for example — relies on assumptions that can end up having meaningful effects, he says.
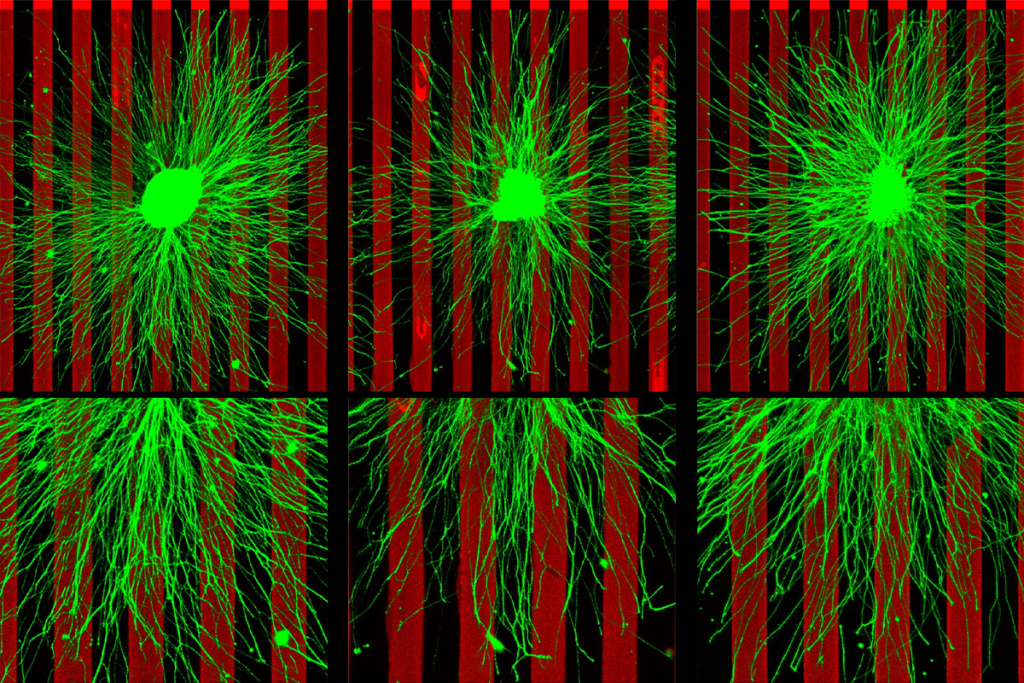
Similarly, different methods researchers use to trace white matter may alter whether two tracts running close to each other are interpreted as crossing like an ‘X’ or just ‘kissing’ and then bending away from each other, says Ruth Carper, research associate professor of psychology at San Diego State University in California. Choices about how to define the border between gray and white matter in the cortex can also skew results.
To address these sorts of problems, the Organization for Human Brain Mapping (OHBM) put out recommendations for analyzing MRI and fMRI data in 2016 and added guidelines for electroencephalography and magnetoencephalography, which are similarly used to collect data from living brains, the following year. The organization is working on an update to the recommendations.
The idea behind the guidelines is to help researchers make better choices throughout their analysis pipeline, says John Darrell Van Horn, professor of psychology and data sciences at the University of Virginia and co-chair of the OHBM’s best-practices committee.
The guidelines also encourage “everybody to be transparent about [the analyses] they’ve performed, so that it’s not a mystery,” Van Horn says.
In addition to improving replicability, more transparency makes it easier for researchers to figure out if one analysis pipeline is better than another, says Tonya White, associate professor of child and adolescent psychiatry at Erasmus University in Rotterdam, the Netherlands, who also worked on the OHBM guidelines.
“If you come up with a new algorithm that you’re saying is better, but you don’t compare it to the old method — how do you really know it’s better?” she says.
Holding up:
For many scientists, better image collection and analysis is the only way forward. Despite the challenges involved — and reports that researchers are turning away from brain imaging — for certain lines of inquiry, there is no substitute.
“You can’t ask about language in a rat” or about brain changes over time using postmortem tissue, Carper says.
Plenty of imaging findings do hold up over time: Certain brain areas activate in response to the same stimuli across all people; people do have individual patterns of connectivity; and scans can predict important categories of behavior, such as executive function and social abilities, Pelphrey says. And when results are inconsistent, as in fMRI studies of reward processing in autism, some researchers have begun implementing an algorithm that helps home in on brain regions that are consistently activated in response to the task.
“There’s lots of stuff that [has been] replicated from lab to lab really nicely and has led to constraints on theories of how the mind works, by virtue of knowing how the brain works,” Pelphrey says. “I don’t think any of that has gone away, just because we know that we have to be very careful about the predictions we make from scan to scan within an individual.”
As scientists get better at collecting and analyzing imaging data, they can also improve on their interpretations of it, built on a clearer understanding of the technology they’re using, Carper says. For fMRI, for example, most tasks were developed to identify brain areas that generally respond to a given stimulus, not to probe individual differences. And certain types of naturalistic stimuli may produce more reliable results, Elliott and his colleagues reported in a review this month in Trends in Cognitive Sciences.
The new review outlines other strategies for sidestepping the fMRI reliability problems the team pointed out last year. Strategies include running longer scans to collect more data, developing better ways to model the noise within a scan and taking advantage of newer technologies that offer better isolation of markers of neural activity.
Elliott did turn to Twitter to promote the new work, he says, but he’s not interested in another social-media controversy. He’s just excited to hear what people think about the paper itself.
“I want to reflect on [the ideas] in a better, longer form than a tweet.”