Insights for autism from Angelman syndrome
Deletions or duplications of the UBE3A gene lead to both Angelman syndrome and some cases of autism, respectively. Studying the effects of altered gene dosage in this region will provide insights into brain defects and suggest targets for therapies for both disorders, says expert Benjamin Philpot.
The word ‘autism’ has unfortunately entered our common lexicon, but few people have heard of Angelman syndrome, a closely linked disorder that is also accompanied by deficits in cognition and language. Autism is frequently diagnosed in individuals with Angelman syndrome1,2.
Angelman syndrome also shares a common genetic basis with some forms of autism: disruption in the number of copies of the UBE3A gene. Studying the effects of altered UBE3A gene dosage is likely to provide insight into brain defects associated with Angelman syndrome and autism, and to suggest targets for therapies.
With the exception of sex chromosome-linked genes, one copy of every gene in the body is inherited from each of an individual’s parents. People usually have both maternally- and paternally-inherited copies of these alleles, but there are some genes — termed imprinted genes — for which only one copy is functional.
UBE3A is one example of an imprinted gene. Only the allele inherited from the mother is functional in neurons, whereas the paternal allele is silenced. This silencing is mediated by epigenetics — a process in which chemical modifications to DNA alter gene expression without changing the primary sequence. Thus, expression of UBE3A in neurons is entirely dependent upon expression from the maternally-inherited allele. The most common cause of Angelman syndrome is a deletion of the maternal chromosomal region 15q11-13, which includes the UBE3A gene. Mutations in the maternal UBE3A gene alone are sufficient to cause Angelman syndrome.
Molecular intersection:
Although multiple genes are likely to contribute to most cases of autism, one of the most genetically identifiable forms of autism is a maternal duplication of the same 15q11-13 region3,4,5. The fact that maternal, but not paternal, duplications of this region are associated with autism implicates the UBE3A gene, because UBE3A is the only paternally imprinted gene in this region.
Although speculative and in need of rigorous testing, these observations suggest that multiple copies of the UBE3A gene can lead to 15q-linked autism. A lack of a functional UBE3A gene copy in neurons, of course, results in Angelman syndrome.
Considering the potential importance of UBE3A gene dosage in both Angelman syndrome and 15q-linked autism, there is increased interest in studying the role of UBE3A in neural functions. Surprisingly little is known about UBE3A, but this is beginning to change.
We know that UBE3A is a ubiquitin E3 ligase that is involved in the degradation of proteins via the ubiquitin-proteasome pathway. Because point mutations that eliminate UBE3A ubiquitin ligase activity are sufficient to cause Angelman syndrome, the most likely explanation for Angelman syndrome — and for 15q-linked autism — is accumulation or enhanced degradation of UBE3A targets.
There are potentially many UBE3A targets and these may be relevant to multiple signaling pathways across different brain regions and periods of brain development. In this way, inappropriate levels of UBE3A may account for the multiple hits that are believed to underlie most cases of autism. Accordingly, a clear research goal is to identify UBE3A substrates in the brain that contribute to neural dysfunction, and the resulting cognitive and behavioral deficits.
Researchers are beginning to identify these brain-specific UBE3A targets6. Last year, Michael Greenberg‘s laboratory at Harvard Medical School identified two protein targets of UBE3A, called ARC and EPHEXIN-5, that affect synaptic receptors — proteins involved in neurotransmission7,8.
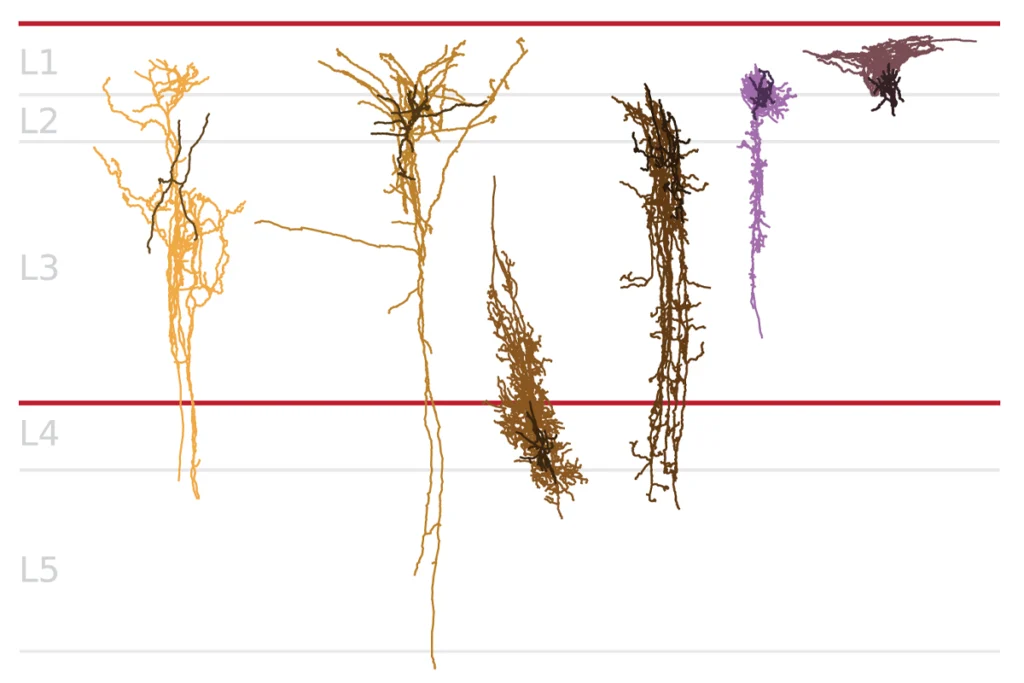
Combined with the subtle changes in brain morphology seen in people with Angelman syndrome and autism, these observations support the idea that both are largely disorders of the synapse, the junction between neurons9.
Postmortem studies of Angelman syndrome have revealed brain region-specific reductions in the density of dendritic spines — the sites of connections between neurons. Changes in synapse density and morphology may also be common features of autism spectrum disorders10.
Common circuits:
By identifying brain regions with impaired synapses caused by inappropriate UBE3A levels, we aim to understand how disrupted circuits contribute to both Angelman syndrome and autism spectrum disorders.
Research published in the past few years indicates that lack of the UBE3A protein prevents brain circuits from strengthening in response to experiences in the mouse sensory neocortex, a brain region involved in regulating sensory perceptions11,12. For example, in mice, UBE3A is required for ocular dominance plasticity — a well-defined model in which brain circuits are refined in response to experience during a critical period of brain development.
Changes in the ability to modify brain circuits in response to experience could underlie the sensory processing and cognitive deficits associated with Angelman syndrome. Whether excessive UBE3A, as seen in 15q-linked autism, also results in altered circuit plasticity remains to be tested. We suggest that the circuits affected in Angelman syndrome are similar to those disrupted in 15q-linked autism and perhaps other autism spectrum disorders as well.
Because Angelman syndrome and 15q-linked autism result from a clear genetic cause, these disorders are comparatively more tractable for therapeutic intervention than are other forms of autism. By studying the mechanisms of UBE3A gene imprinting and the role of UBE3A in circuit development, our goal is to help pave the way for novel therapies for both disorders.
Behavioral interventions can partially restore synapse and circuit plasticity in Angelman syndrome mice12. This suggests that the mechanisms underlying synaptic plasticity remain intact in the absence of UBE3A. If the same applies to autism, then behavioral or pharmacological therapies have the potential to restore function.
The identification of pathways relevant to synapse dysfunction in Angelman syndrome and 15q-linked autism will largely be based on the identification of brain-specific UBE3A targets. It is likely that UBE3A substrates accumulate to inappropriately high levels in Angelman syndrome but are degraded to inappropriately low levels in 15q-linked autism. Pharmacological approaches could regulate key UBE3A substrates and the pathways through which they signal.
Because UBE3A targets may be too numerous to address individually, manipulating UBE3A expression levels directly is potentially a more promising therapeutic strategy. Indeed, our laboratory, together with our collaborators Mark Zylka and Bryan Roth, has been awarded a grant from the Simons Foundation, SFARI.org’s parent organization, to identify ways to manipulate UBE3A levels as a treatment for autism spectrum disorders.
Benjamin Philpot is associate professor of cell and molecular physiology at the University of North Carolina School of Medicine. Angela Mabb and Matthew Judson are postdoctoral fellows in his lab.
References:
- Steffenburg S. et al. Pediatr. Neurol. 14, 131-136 (1996) PubMed
- Peters S.U. et al. Clin. Genet. 66, 530-536 (2004) PubMed
- Schanen N.C. Hum. Mol. Genet. 15, 138-150 (2006) PubMed
- Cook E.H. Jr. et al. Am. J. Hum. Genet. 60, 928-934 (1997) PubMed
- Glessner J.T. et al. Nature 459, 569-573 (2009) PubMed
- Scheiffele P. and A.A. Beg Nature 468, 907-908 (2010) PubMed
- Greer P.L. et al. Cell 140, 704-716 (2010) PubMed
- Margolis S.S. et al. Cell 143, 442-455 (2010) PubMed
- Zoghbi H.Y. Science 302, 826-830 (2003) PubMed
- Betancur C. et al. Trends Neurosci. 32, 402-412 (2009) PubMed
- Sato M. and M.P. Stryker Proc. Natl. Acad. Sci. U. S. A. 107, 5611-5616 (2010) PubMed
- Yashiro K. et al. Nat. Neurosci. 12, 777-783 (2009) PubMed
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question