Method reads brain cell signatures to reveal risky mutations
A new approach ranks genes’ ties to autism based on their expression patterns in different types of brain cells.
A new approach ranks genes’ ties to autism based on their expression patterns in different types of brain cells1. The method could help researchers predict the effects of rare, spontaneous mutations in autism.
Scientists have looked for a ‘molecular signature’ for autism in genetic and other molecular features that people with autism share. The new method gleans a signature from gene expression profiles in cell types such as neurons and astrocytes.
Cell-specific patterns may reveal differences in gene regulation that aren’t apparent in analysis of brain tissue as a whole. These differences may point to molecular pathways that are altered in autism.
Researchers used published data from mice to compile a list of 15,951 genes expressed in 24 types of brain cells. All of the genes are also expressed in people.
The team also used four sequencing studies to list 145 genes that have been shown to carry spontaneous mutations in people with autism or their unaffected siblings. Of these, 112 genes showed up in people with autism alone.
Drawing from two types of information in this way should limit spurious associations that may turn up in one or the other. The work appeared 5 December in Human Mutation.
Settling the score:
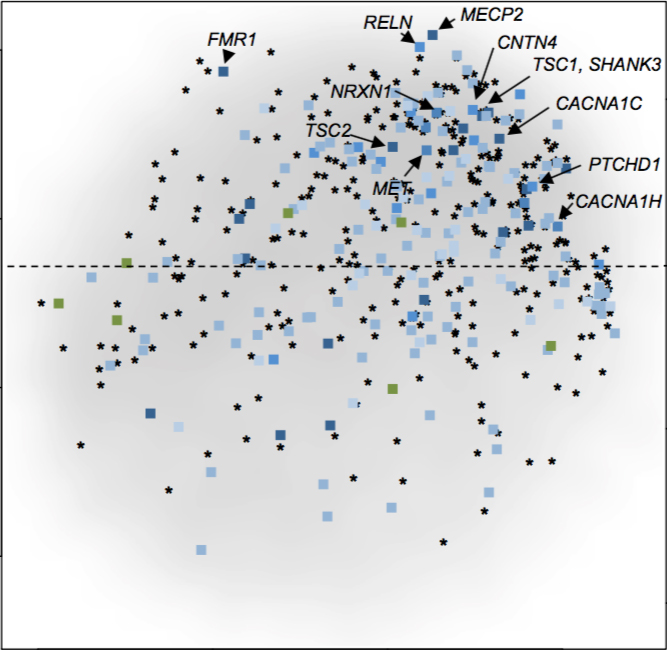
Analyzing the data from both sources, the researchers generated a score that reflects each gene’s likely connection to autism.
They built the scoring method using data published prior to 2013 so they could test it with newer data. The team reanalyzed and scored 611 genes from two later sequencing studies and 483 genes from a database of ranked autism genes.
The method produced high scores for genes that the database ranks as strongly linked to autism and low scores for those with lower status, suggesting that it is accurate.
Gene expression patterns revealed that mutations in 117 of the highest-scoring genes typically destroy the gene’s function. Many of these genes are involved in regulating the expression of other genes.
High-scoring autism risk genes tend to be switched off or tamped down in certain types of neurons. Such cell-specific findings could help scientists understand how certain mutations contribute to autism features.
- Zhang C. et al. Hum. Mutat. 38, 204-215 (2016) PubMed
Recommended reading

Building an autism research registry: Q&A with Tony Charman
CNTNAP2 variants; trait trajectories; sensory reactivity
Brain organoid size matches intensity of social problems in autistic people
Explore more from The Transmitter
Cerebellar circuit may convert expected pain relief into real thing