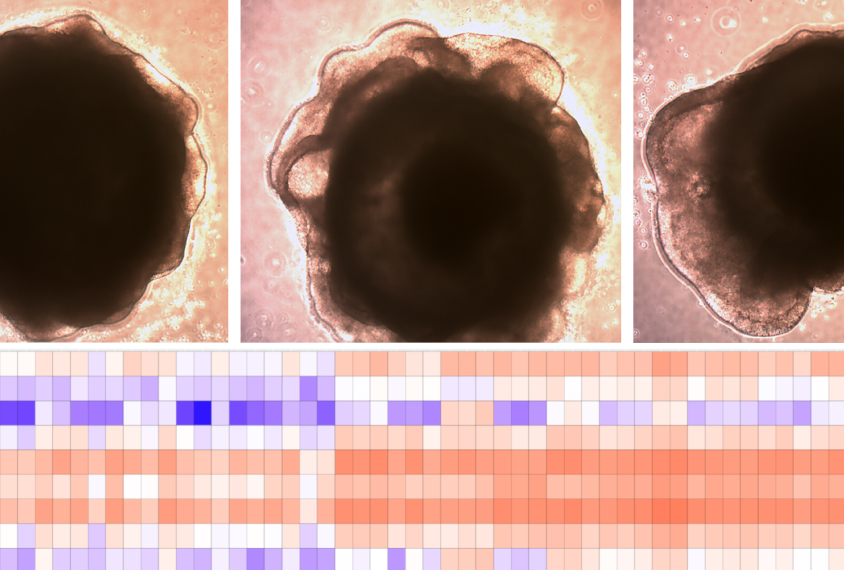
A new method for analyzing gene expression levels in human brain organoids — spheres of cultured brain cells — enables researchers to determine whether DNA mutations linked to autism affect some cell types more than others.
Researchers used the method, called Orgo-Seq, to show that deletions of a chromosomal region called 16p11.2 increase the proportion of immature neurons and neural progenitor cells — early precursors of neurons and glia — in brain organoids. The method also revealed genes in the region that may drive those changes.
16p11.2 deletions can result in neurodevelopmental conditions, including autism. The new technique allows researchers to “go after cell types implicated by different genes that have been associated with autism,” says lead investigator Elaine Lim, assistant professor of molecular, cellular and cancer biology at the University of Massachusetts in Worcester.
Lim and her colleagues grew organoids from skin cells taken from 9 people with 16p11.2 deletions, 4 people with a duplication in chromosomal region 15q11-13, which is also associated with autism, and 12 people without such deletions or duplications.
Typical brain organoids contain at least 10 distinct categories of cell types, according to previous studies that distinguished the categories by their gene expression patterns, one cell at a time. But such single-cell analyses are expensive, and the patterns they detect can vary from organoid to organoid — even between those derived from the same person.
L
im and her colleagues developed a strategy to infer the expression patterns in individual cells of the organoids without having to isolate and analyze single cells. They measured the gene expression levels across entire organoids in bulk and then focused on the genes specific to 10 cell-type categories from a 2017 study and 11 from a 2020 study. They reasoned that differences in the expression of these genes between control and mutant organoids would reflect differences in the proportion of the related cell types.The analysis revealed an increase in immature neurons and neuron precursors in 16p11.2 organoids, but it did not turn up any cell-type differences in 15q11-13 organoids.
Lim’s team then focused on the 29 genes within the 16p11.2 region. They searched for these genes in gene expression databases to identify those that are expressed during the same time in brain development as the cell-type-specific genes that are altered in the 16p11.2 organoids. Their search turned up three strongly linked genes — YPEL3, KCTD13 and INO80E. They conclude that these genes drive the cell-type differences in the organoids. Some previous studies have implicated KCTD13 in the brain enlargement seen in zebrafish missing 16p11.2.
The team reported their findings in June in Nature Communications.
The approach represents “an important piece” in the quest to identify the developmental origins of autism, says Xin Jin, assistant professor of neuroscience at Scripps Research in San Diego, California, who was not involved in the study. Although the technique was developed for use in organoids, it could be applied to postmortem brain tissue to determine whether similar changes happen in the brain, Jin and Lim say.
Lim cautions that not all autism-linked mutations can be tackled using her team’s approach, which is best suited for mutations that affect the balance of cell-type proportions.