Common genetic variants shape the structure of the cortex
A genome-wide association study lays a foundation for deeper investigation of these variants in neurodevelopmental conditions.
Some 4,349 regions in the genome seem to govern key physical characteristics in the cerebral cortex, according to a new genome-wide association study (GWAS). The research included genetic and MRI data from more than 36,000 people.
This is the most detailed study of brain phenotypes in a sample this large to date, says lead investigator Richard Bethlehem, director of neuroimaging at the Autism Research Centre at the University of Cambridge in England.
Many of the genetic areas, or loci, identified overlap with those flagged in other studies of brain structure, such as work by the Enhancing Neuro Imaging Genetics through Meta Analysis (ENIGMA) consortium. But ENIGMA uses neuroimaging data from many sites with varied scanning protocols, whereas the new study draws from just two datasets, making the imaging data more homogeneous, Bethlehem says.
The new statistical analyses, which are publicly available, “give people a map” to investigate the connection between genes associated with neurodevelopmental conditions and those that drive brain growth, Bethlehem adds.
“The most impressive part was actually the comprehensiveness of this analysis,” says Juergen Dukart, professor of digital biomarkers at Heinrich-Heine-Universität in Düsseldorf, Germany, and group leader of biomarker development at Forschungszentrum Jülich, who was not involved in the new work. “It just gives a very, very convincing story, starting from the genes to the brain phenotypes.”
The findings were published in August in Nature Genetics.
B
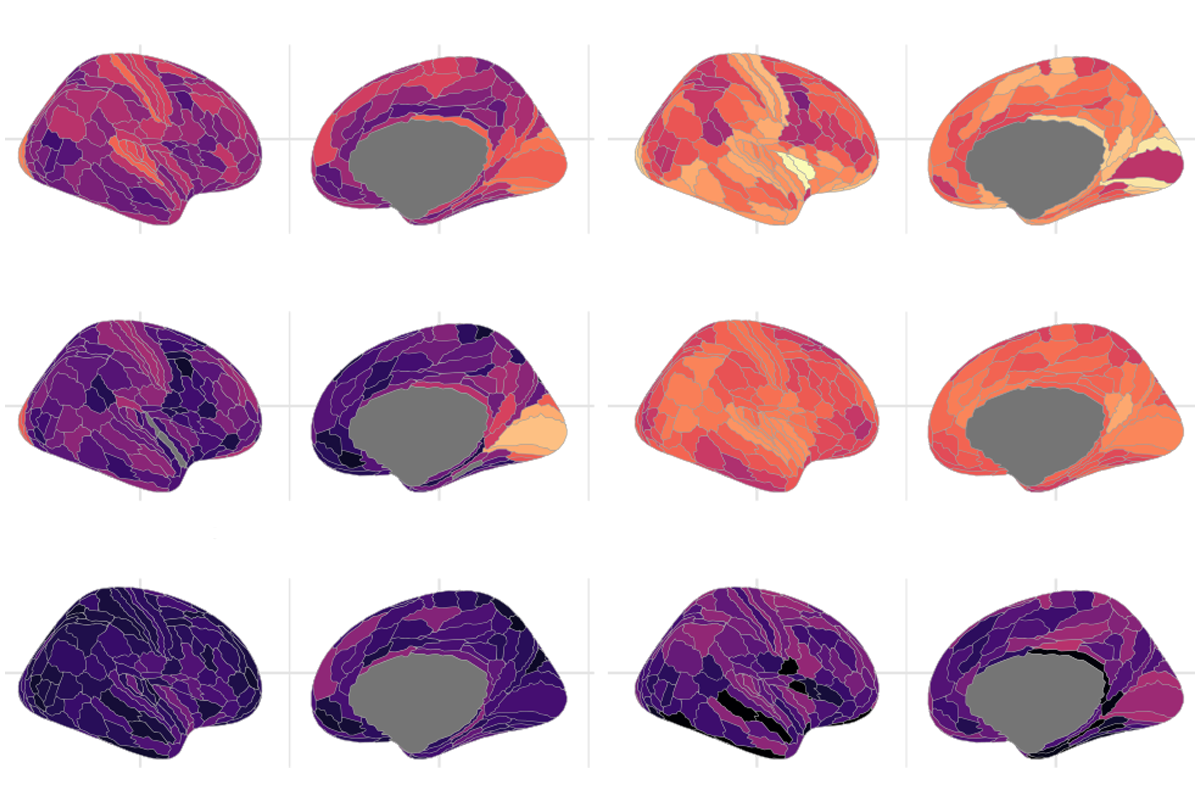
The genetic signatures for 12 of the phenotypes (all except cortical thickness) resembled each other so closely that the researchers clustered them into four broader characteristics: neurite density and orientation, cortical expansion, curvature, and water diffusion. This finding indicates the 13 parameters “might be different ways of measuring the same biological construct,” Bethlehem says. “I wasn’t aware of anyone who had shown that at this level.”
In total, 4,349 loci cropped up as significantly related to the four aspects of cortical structure. The team then homed in on 40 candidate genes for brain structure.
Past research has identified four of these candidates in groups of people with atypical brain size and neurodevelopmental conditions. The gene KANSL1, for example, is associated with Koolen-de Vries syndrome, which is characterized by global developmental delays and, in more than half of cases, microcephaly. Another candidate gene, HMGA2, is associated with macrocephaly and Russell-Silver syndrome, which is characterized by learning problems.
The fact that these genes — previously discovered via rare variants — were flagged in the new study via the presence of common variants is “new and interesting,” says Jason Stein, associate professor of genetics at the University of North Carolina at Chapel Hill and co-founder of ENIGMA. “It’s usually thought that developmental disorders are mainly driven by rare genetic variation.”
Stein, who was not involved in the new research, adds that “this is similar to findings from other studies that [show] that it’s not just rare variation that changes phenotypes; it’s a combination of common and rare variation.”
T
In addition, autism can lead to a wide variety of brain changes, so genes associated with brain development may not give a clear signal in groups of people with autism, Bethlehem and his colleagues say.
Bethlehem’s team is now extending their findings to look for any overlap between the genes tied to schizophrenia and the genes their approach has associated with brain features linked to the condition. With larger GWAS datasets, “I think you can have a similar approach for autism,” Bethlehem says.
Future research could also focus on the loci flagged as important for specific brain regions. Those genes could guide animal studies or research in subgroups of people with autism or neurodevelopmental conditions who show differences in those brain regions. Such an approach could lead to identifying biomarkers for these conditions, Dukart says.
By studying multiple measurements from people across the lifespan, researchers could also use the new findings to better understand how specific genes lead to certain phenotypes, Bethlehem says. How environmental factors affect this trajectory is especially important to investigate: Depending on the condition involved, the environment can lead to “quite a significant chunk of individual variability, and we don’t really understand what it’s driven by,” he says.
Recommended reading
New ‘decoder’ tool translates functional neuroimaging terms across labs
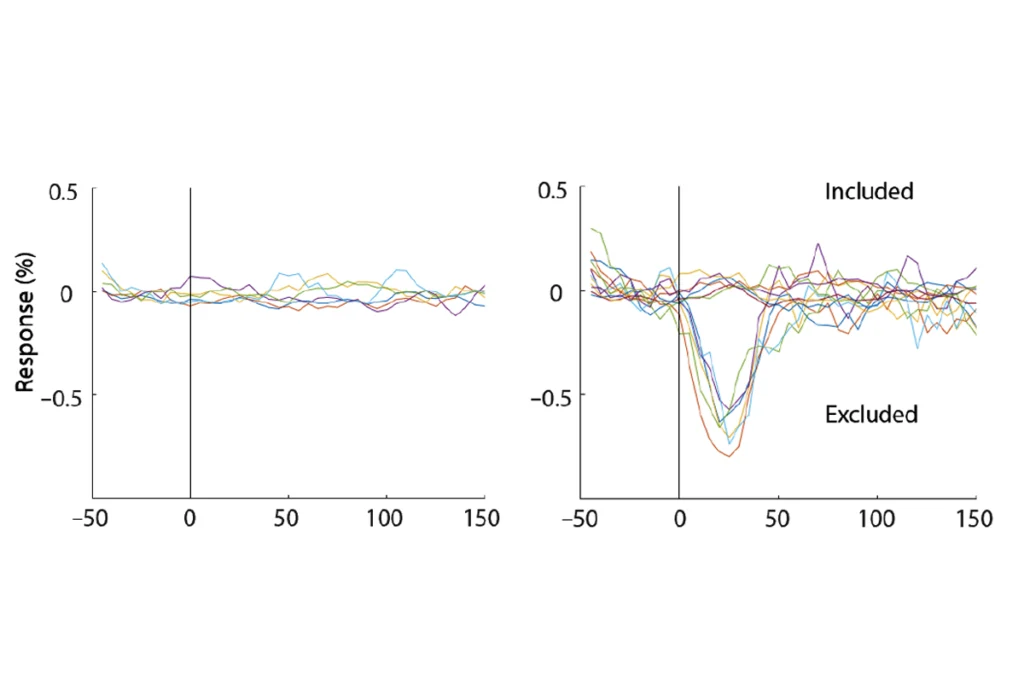
Two studies fail to replicate ‘holy grail’ DIANA fMRI method for detecting neural activity

Name this network: Addressing huge inconsistencies across studies
Explore more from The Transmitter

Building an autism research registry: Q&A with Tony Charman