Neurons exposed to valproic acid (VPA), an epilepsy drug that, when taken during pregnancy or breastfeeding, increases a child’s chances of autism, produce atypical levels of RNA isoforms, according to a new study. Several of these alternate RNA versions are encoded by genes tied to autism, the study also shows.
VPA is known to unravel chromatin, the coiled complex of DNA and proteins in a cell’s nucleus. The new findings suggest that environmental perturbations that alter chromatin structure can influence not only gene expression but also how a gene’s coding regions are shuffled to produce several versions of a protein. This process, called alternative splicing, has been implicated in autism.
Altered splicing patterns may also result from mutations in chromatin-regulating genes, some of which are linked to autism and other neurodevelopmental conditions, says Sofia Lizarraga, assistant professor of molecular biology, cell biology and biochemistry at Brown University in Providence, Rhode Island, who co-led the new research. For example, dampening the levels of the autism-linked chromatin regulator CHD8 alters splicing in neurons, a December study showed.
“Understanding these mechanisms is going to be important to be able to develop therapeutic strategies in the future,” Lizarraga says.
L
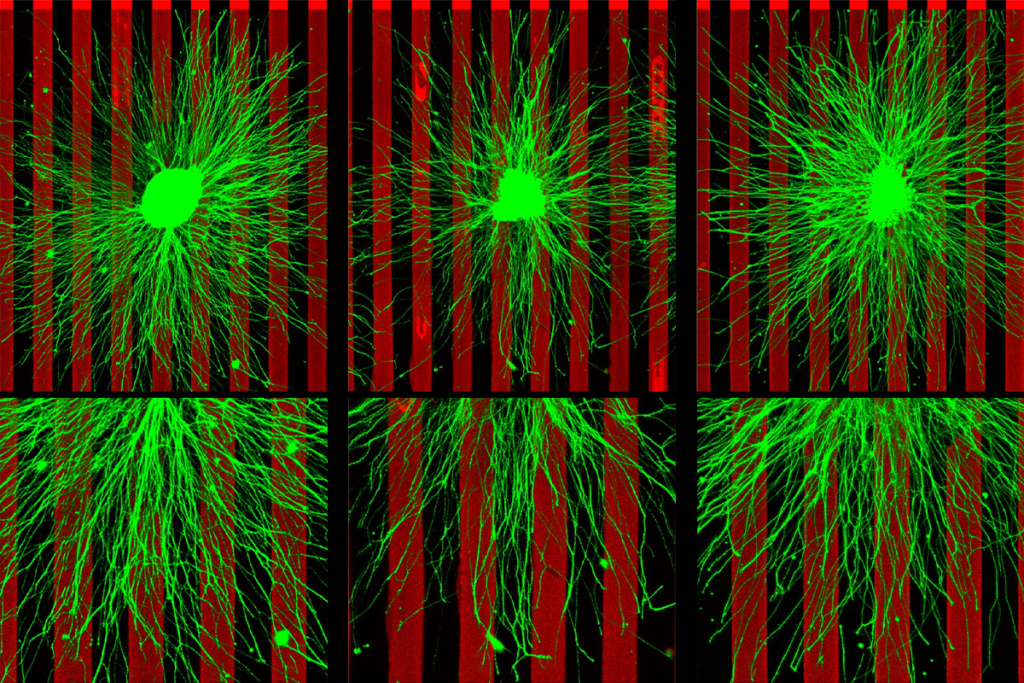
izarraga and her colleagues reprogrammed skin cells from a neurotypical person to differentiate into cortical neurons that send excitatory signals. After 24 hours of exposure to VPA, the neurons showed altered expression levels of more than 6,000 genes compared with control neurons. Several of the genes with dampened expression help regulate RNA processing, suggesting that VPA can influence the RNA isoforms that neurons produce. Indeed, the treated neurons produced different levels of RNA isoforms from about 3,800 genes, the researchers found.Those genes tend to be involved in transcription and chromatin regulation, whereas differentially expressed genes are associated with synaptic structure and function. About 25 percent of these two classes of genes overlap with autism-linked genes, including CHD2 and SETD5, the researchers found.
An analysis of SETD5 transcripts showed that treated neurons had reduced levels of an isoform that encodes the full-length protein and increased levels of one noncoding isoform. Similarly, treated neurons showed decreased levels of two protein-coding isoforms of the gene SMARCA4, which has recently been tied to autism.
“We saw several examples in which the main coding isoform was down, and the noncoding or truncated isoforms were up,” Lizarraga says. “That could really be affecting the biology [of neurons].”
The findings appeared in Human Molecular Genetics in January.
V
PA alters chromatin structure by inhibiting a class of enzymes called histone deacetylases (HDACs), which remove acetyl tags from the proteins around which DNA wraps.Lizarraga and her colleagues propose three models through which VPA-induced inhibition of HDACs could lead to altered splicing patterns: by changing the rate at which the transcription machinery travels along DNA; by directly hindering the interaction between the enzymes and splicing factors, which piece together different RNA segments; or by preventing histones from recruiting those factors to chromatin.
Testing these models and assessing in vivo how VPA affects different types of neurons at different developmental stages would be important, says Maria Chahrour, associate professor of neuroscience and human growth and development at the University of Texas Southwestern Medical Center in Dallas, who was not involved in the work.
“It would be also interesting to see if [altered splicing] is happening in genetic autism models where chromatin remodelers are disrupted,” Chahrour says.
T
he researchers characterized RNA transcripts using a sequencing technology that reads a few hundred base pairs at a time. Newer methods that sequence an entire transcript in one go may reveal additional isoforms that are produced at different levels in the VPA-exposed neurons, says Jonathan Mill, professor of epigenetics at the University of Exeter in the United Kingdom, who wasn’t involved in the study. “This [work] is probably highlighting the tip of the iceberg, and it does suggest that there might be many more of these events to be found,” he says.Regardless, the findings add to mounting evidence that specific RNA isoforms contribute to some neurodevelopmental conditions — offering clues for potential treatment strategies.
“There’s huge excitement around things like antisense oligonucleotides, for instance, that can target the RNA molecule and silence specific transcripts,” Mill says. If researchers identify specific RNA variants that are differentially produced as a result of an environmental exposure or a genetic mutation tied to autism, these types of drugs may help restore typical levels of those isoforms, he says.
Next, the researchers plan to look at other brain cell types, using single-cell analysis of RNA transcripts, says study co-lead investigator Ece Gamsiz Uzun, assistant professor of pathology and laboratory medicine at Brown University.
“Can we see these mechanisms in astrocytes or oligodendrocytes and other cell types that are important for brain development? Can we rescue some of these alterations by re-expressing some of the proteins that are downregulated? We’d like to get at it in a more functional way,” she says.