Mutated DNA loops make strange neighbors
Too many or too few copies of a segment of chromosome 16 alters the three-dimensional organization of DNA, and affects hundreds of related genes.
Too many or too few copies of a segment of chromosome 16 alters the three-dimensional (3D) organization of DNA, and affects hundreds of related genes, suggests a new study1.
Within a cell, DNA coils around proteins to form a 3D structure called chromatin. The coil may bring distant genes close together, or pull nearby genes apart.
The new study suggests that genes that cluster together in 3D space play similar roles. Changes to one segment may jumble this structure and tweak the expression of many genes.
The study focused on deletions or duplications of a 600-kilobase stretch called 16p11.2, which sometimes lead to autism. These variants alter the location and expression of dozens of genes in other regions, the study found.
“The human genome is probably not organized in a totally random manner,” says lead researcher Alexandre Reymond, director of the Center for Integrative Genomics at the University de Lausanne in Switzerland. “Here we have multiple genes that are extremely packed in space, and many of those are playing a role in the same pathway.”
Reymond’s team presented preliminary results at the 2014 annual meeting of the American Society of Human Genetics in San Diego. They published the findings in May in Molecular Psychiatry.
The findings are specific to copy number variations (CNVs) — deletions or duplications of long segments of DNA — but may also extend to mutations of other types, says Dan Arking, associate professor of genetics at John Hopkins University in Baltimore, who was not involved in the study.
“Looking more broadly at regulation through three-dimensional chromosomal structure and interaction is likely to be important in autism and complex disease in general,” he says.
Mirror effect:
Reymond and his colleagues first noticed the importance of chromatin structure when they explored genes in a 220-kilobase segment of chromosome 16. These genes are located more than 800 kilobases away from those in the better-known 600-kilobase region.
They discovered that, despite the distance, 88 people with deletions and 49 with duplications of the smaller region have features that are strikingly similar to those seen in people with CNVs in the longer stretch: Deletion of either region leads to obesity and a large head, whereas duplication is tied to low body weight and a small head. And roughly 25 percent of people with a deletion or duplication of either region have autism.
These similarities may be due to the fact that in the 3D structure, genes in both regions are close together, the researchers say.
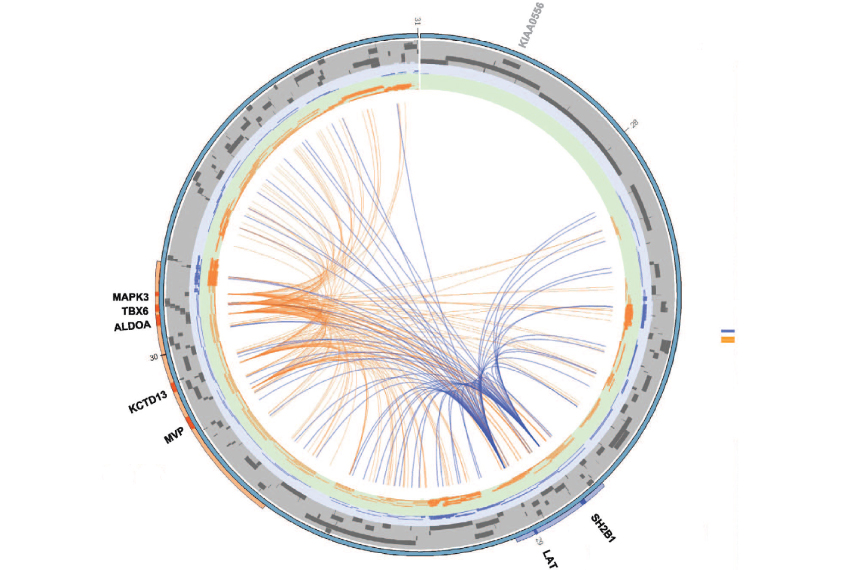
They used a method that binds together chromosomal regions in close proximity and then sequences across the junctions. MAPK3 and KCTD13 — two genes in the longer segment that have strong ties to autism features — cluster near genes in the smaller region, they found.
They then repeated this experiment in cells from two people with a deletion and two with a duplication of the smaller region. The CNVs dramatically shifted the location of 1,193 genes — many of which are linked to autism, body mass index or head circumference.
Discovery hub:
The same researchers showed last year that 125 of these genes have altered expression in people with duplications or deletions of the larger region. This list includes 12 autism candidates, 5 of which are linked to head size.
Together, the two studies suggest that genes that work together — and may need to be turned on or off at the same time — also cluster together in physical space, says Reymond. “Maybe those [chromatin] contacts will help us find genes that are in the same pathway and playing the same role,” he says.
To test this theory, the researchers focused on two genes in a section of chromosome 2 that cluster near the larger 16p11.2 region. The two genes move away when the region is duplicated or deleted.
The researchers then looked at 35 people with CNVs in the chromosome 2 segment. They found that 26 people with a deletion of that section have a low body mass index and small head, and 9 people with duplications show the opposite pattern.
These features mirror those seen in people with CNVs in the 16p11.2 region. The finding suggests that variants of different segments in the same 3D cluster lead to similar effects.
How neighboring genes from different chromosomes are turned on or off together still needs to be worked out, says Fulai Jin, assistant professor of genetics and genome sciences at Case Western Reserve University in Cleveland, who was not involved in the new work.
It’s unclear whether the results would hold up with other CNVs, or in neurons rather than blood cells used in the study, Jin adds. Reymond’s team is testing whether the findings hold true in mouse neurons.
References:
- 1: Loviglio M.N. et al. Mol. Psychiatry Epub ahead of print (2016) PubMed
Explore more from The Transmitter

At 25, INSAR needs to bring autism scientists together more than ever