Study maps genetic variability in autism brains
The first effort to sequence genes tied to autism in postmortem brain tissue reveals a range of harmful mutations in people with the condition.
The first effort to sequence genes tied to autism in postmortem brain tissue reveals a range of harmful mutations in people with the condition.
The findings, reported 2 December in Neuron, underscore the tremendous diversity of genes that can contribute to autism. They also lend support to studies that flagged the same genes in more accessible tissues, such as saliva or blood1.
“A significant proportion of brains in the postmortem autism collections have identifiable, deleterious mutations that are likely to be causative,” says lead investigator Christopher Walsh, professor of neurology at Harvard Medical School. The results allow researchers to compare the effects of the different mutations on the brain, Walsh says.
Sequencing studies have shown that most of the genetic risk for autism is inherited, whether through rare mutations or common variants. These studies have also uncovered hundreds of autism-linked genes that carry de novo mutations, which arise spontaneously in a mother’s egg or a father’s sperm.
The new work provides important confirmation that autism-linked mutations seen in blood and saliva also appear in the brain. It also implicates an additional type of genetic risk in autism: so-called ‘somatic mutations.’ Unlike de novo mutations, which occur throughout the body, somatic mutations arise spontaneously sometime after conception and appear in only some cells.
“It certainly looks like somatic mutation is not the leading cause of autism by any stretch, but there’s a chance it plays an important role,” says Stephan Sanders, assistant professor of psychiatry at the University of California, San Francisco, who was not involved in the work.
Two of the brains used in the study carry somatic mutations in autism-linked genes that show up in specific regions of the brain. Sequencing more postmortem brain samples may reveal brain regions that are important in autism, Sanders says.
Going deep:
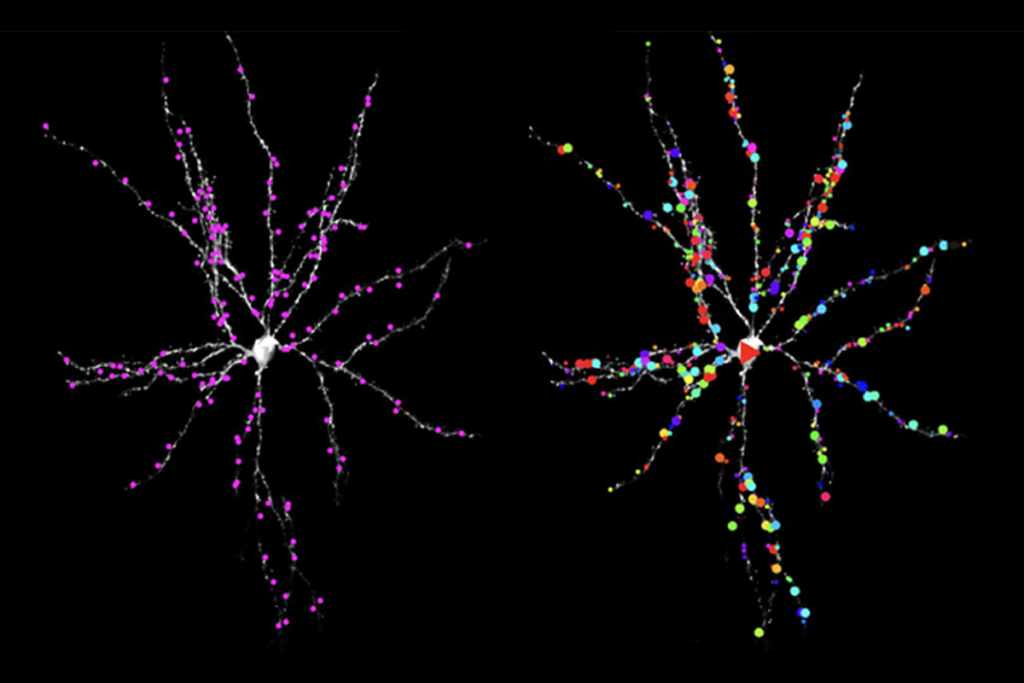
Walsh’s team studied postmortem brain tissue from 55 people with autism and 50 controls, obtained from two major repositories of such tissue: Autism BrainNet and NeuroBioBank. The researchers looked at samples from two brain regions that are implicated in autism: the cerebellum, which helps to control and coordinate movements, and the frontal cortex, which is involved in cognition, sensory processing and social behavior.
They analyzed 78 genes strongly linked to autism, sequencing each gene nearly 550 times per sample to distinguish real mutations from random sequencing errors. This ‘deep-sequencing’ strategy allowed the researchers to spot somatic mutations, which are difficult to detect using conventional sequencing because they appear in only a fraction of the cells.
Brain tissue from people with autism is more likely to carry a harmful mutation than control brain tissue, the researchers found. Of the 55 autism samples, 16 carry harmful mutations, compared with 5 of 50 control samples.
The team uncovered 34 damaging mutations across 25 genes in the autism samples. One gene, SCN2A — one of the strongest autism candidate genes identified to date — is mutated in four of the autism brains. Three other genes — ARID1B, SCN1A and SETD2 — carry various mutations in two autism brains each.
Because the researchers didn’t have access to DNA sequences from the parents or siblings of the donors, they cannot tell whether the mutations are inherited or spontaneous. But the excess number of mutations in the autism samples suggests that, either way, these mutations are likely to contribute to the condition.
Patchwork puzzle:
One of the study’s most surprising findings is that two of the brains carry distinct somatic mutations.
“Even by just looking at a handful of genes and samples, they’re actually starting to find [somatic] mutations,” says Brian O’Roak, assistant professor of molecular and medical genetics at Oregon Health and Science University in Portland, who was not involved in the study. “I think this tells us we should keep looking.”
One of the somatic mutations is present in the frontal cortex but not the cerebellum. The other appears in both regions, but only in some cells in each region. The distinct distribution of the two mutations suggests that they arose at different times in prenatal development, says Eric Courchesne, professor of neuroscience at the University of California, San Diego, who was not involved in the work.
It also highlights the difficulty of identifying somatic mutations, which might occur only in subsets of brain regions. “I think what they found might be sort of the tip of the iceberg,” Courchesne says.
Walsh and his colleagues examined the medical records of the donors whose brain samples yielded mutations. In six cases, the combination of the symptom profile with the newly identified gene mutation suggests that the individual may have had an autism-linked syndrome. For instance, two individuals who had SCN1A mutations had experienced seizures starting in infancy, suggesting that they may have had Dravet syndrome.
The analyses are limited by the number of postmortem brains used. A larger pool of specimens would allow researchers to match a given genetic glitch with a pattern of cellular alterations, Courchesne says.
“It would be fascinating to compare and contrast a variety of experiments examining cellular size, neuron types and other characteristics in these cases,” Courchesne says. “There’s a host of really important studies to do.”
References:
- D’Gama A.M. et al. Neuron 88, 910-917 (2015) PubMed
Explore more from The Transmitter