The value of blood cells in autism research
Blood from individuals with autism could help researchers identify biomarkers to diagnose the disorder and learn more about related symptoms, such as gastrointestinal complaints, says molecular biologist Valerie Hu.
-

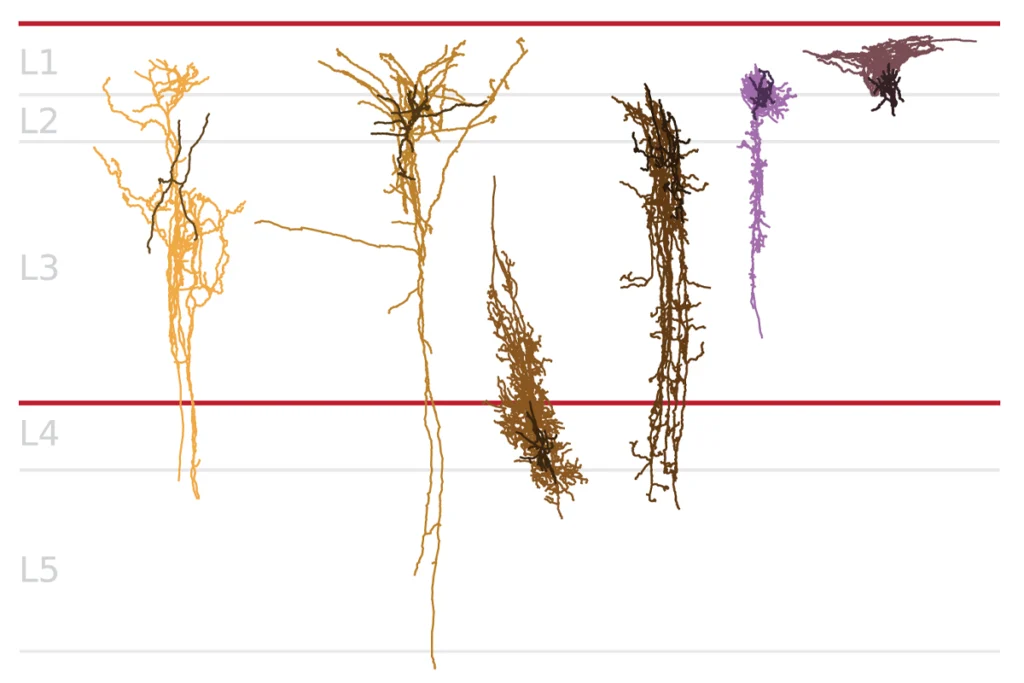
Simple solution: Blood can substitute for brain tissue in some studies of gene expression in autism.
- Simple solution: Blood can substitute for brain tissue in some studies of gene expression in autism.
Autism spectrum disorders result in striking core deficits, such as impaired communication and social skills, and repetitive stereotyped behaviors. However, many individuals with autism also have other symptoms, such as gastrointestinal complaints, immune system complications and sleep disturbances. Identifying biomarkers for autism using blood cells could be one way to diagnose the disorder and learn more about these related symptoms.
The variety of symptoms across individuals with autism and the fact that multiple organs are involved suggest that autism could be a ‘systems disorder,’ which means that the function of tissues outside of the brain may also be affected. To resolve this question, researchers can study autism using easily accessible tissues, which may reflect abnormalities that are also present in the brain. Identification of consistent biomarkers in cells from peripheral tissues, such as blood, could also allow researchers to develop new tests for earlier and more definitive autism diagnosis.
Lymphoblastoid cell lines are continuously dividing cells derived from blood lymphocytes, white blood cells that play a crucial role in the immune system. The cell lines have been widely accepted as a tool to examine the consequences of genetic mutations on the expression of autism candidate genes. However, over the past five years, several research groups have employed these cell lines as tools to investigate large-scale alterations in gene expression as well as epigenetic changes — which modify gene expression without altering the DNA sequence.
The advantage of lymphoblastoid cells is that they are readily available from cell repositories such as the Autism Genetic Resource Exchange and, more recently, the Simons Simplex Collection. The latter is a database of genetic and clinical information from families that have one child with autism and unaffected parents and siblings, funded by SFARI.org’s parent organization. Together, these repositories have collected blood from thousands of well-characterized individuals who have a carefully validated autism diagnosis. These repositories also have blood from the affected individual’s parents, siblings and extended family, enabling comparisons with control groups and analyses of inheritance.
The main purpose of these collections is to provide a continuous source of genetic material through the immortalization and maintenance of these cultured cells. On the other hand, the use of primary lymphocytes, which are collected directly from individuals diagnosed with autism and their families, offers the advantage of not having been subjected to immortalization procedures, continued passaging and storage, which can affect both gene expression and epigenetics.
The disadvantages of primary cells are that it is time-consuming to recruit and validate sufficient individuals with autism spectrum disorder for the studies, and it is harder to control for the individual’s health or medication status at the time of blood collection. The cells are also not renewable through cell culture.
New insights:
Both immortalized and primary blood cells have offered valuable insight into the genes and biological processes affected by autism. Surprisingly, many of the genes that are expressed differently in the lymphoblastoid cells of individuals with autism compared with controlsare associated with neurological functions. A major caveat when using any peripheral tissues is that the disrupted genes and pathways identified may not be altered in the brain because both gene expression and epigenetic regulation are known to be tissue-specific.
Replicating molecular changes seen in peripheral cells in brain tissue offers further support for the involvement of these genes in the etiology, or at least the neuropathology, of autism. For example, a 2006 study reported the first use of lymphoblastoid cells for investigating differences in gene expression between identical twins of whom only one has autism1. This technique makes use of the fact that twins have the same genome to focus on differences in gene expression.
Interestingly, approximately half of the differentially expressed genes identified in the study are involved in nervous system development and function, including neurite extension and axon guidance (ROBO1 and CHL1) as well as neuronal development and survival (DAPK1 and EGR2). Surprisingly, many of the differentially regulated genes are also involved in inflammation (ASS, ALOX5AP and IL6ST). This finding is consistent with evidence of inflammation and activation of microglia, which are inflamed in the postmortem brains of individuals with autism.
Another study by the same laboratory on 21 sibling pairs in which one sibling has autism further reveals the involvement of steroid hormone metabolism in the disorder2. A 2009 study using lymphoblastoid cells from 87 unrelated individuals, divided by subtype of autism, shows subtype-specific expression of genes. Many of the genes discovered in the original twin study were corroborated by the later studies, suggesting that some of the genes may be useful biomarkers for autism3.
In another study, researchers profiled three age-matched pairs of siblings, in which only one of the two siblings is diagnosed with autism4. This study supported the idea that the immune system is dysfunctional in individuals with autism. Another strategy is to investigate the overlap in gene expression between individuals with fragile X syndrome and those who have a duplication of a region on chromosome 15. Two of the identified genes are also misregulated in the blood cells of individuals with idiopathic, or non-syndromic, autism, the study found5.
Complex regulation:
The studies described above focused on gene expression differences in autism, but lymphoblastoid cells have also been successfully used to identify differences in microRNAs (miRNA), fragments of RNA that interfere with protein production, and DNA methylation, which regulates gene expression by adding a methyl group to DNA.
A 2008 study reported that nine miRNAs are differentially expressed between six individuals with autism and six age- and sex-matched controls6. A separate study in the same year found that several of these miRNAs are expressed differently in postmortem brain tissue from donors with and without autism7.
Another study, from 2010, also reported 48 differentially expressed miRNAs in lymphoblastoid cells from twins and sibling pairs in which only one sibling has autism. Among these miRNAs, 16 function in the brain. The 48 miRNAs also potentially regulate 1,053 genesthat showed differences in expression in the same samples8.
Again employing lymphoblastoid cells from discordant twin and sibling pairs, researchers in a 2010 study demonstrated differences in DNA methylation between cases and controls. Included in these is a gene that regulates the expression of other genes, called retinoic acid-related orphan receptor alpha, or RORA9. There is less RORA protein in the cerebellum and the frontal cortex in postmortem brains from individuals with autism compared with controls, which was found to correlate with the reduction of another protein, aromatase, that controls the levels of male and female hormones in tissues10.
This finding links epigenetic and molecular alterations in lymphoblastoid cells with brain pathology, reinforcing the relevance of this novel candidate gene for autism as well as the usefulness of these cells in revealing meaningful differences in autism. Though not without its caveats, lymphoblastoid cells can provide a valuable tool in studies of autism, especially for biomarker identification.
The ability to detect molecular differences in blood cells or other easily accessible tissues, such as cheek cells, is essential to the use of biomarkers for diagnosing autism.After all, who in their right mind would want to provide brain tissue for diagnostic screening?
Valerie Hu is professor of biochemistry and molecular biology at the George Washington University School of Medicine and Health Sciences in Washington, D.C.
References:
1: Hu, V.W. et al. BMC Genomics 7,118 (2006) PubMed
2: Hu, V.W. et al. PLoS ONE 4, e5775 (2009) PubMed
3: Hu, V.W. et al. Autism Research 2, 78-97 (2009) PubMed
4: Baron, C.A. et al. J. Autism Dev. Disord. 36,973-982 (2006) PubMed
5: Nishimura, Y. et al. Hum. Mol. Genet.16, 1682-1698 (2007) PubMed
6: Talebizadeh, Z. et al. Autism Res. 1, 240-250 (2008) PubMed
7: Abu-Elneel, K. et al. Neurogenetics 9, 153-161 (2008) PubMed
8: Sarachana, T. et al. Genome Medicine 2, 23 (2010) PubMed
9: Nguyen, A. et al. FASEB J. 24, 3036-3051 (2010) PubMed
10: Sarachana, T. et al. PLoS ONE 6, e17116 (2011) PubMed
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question