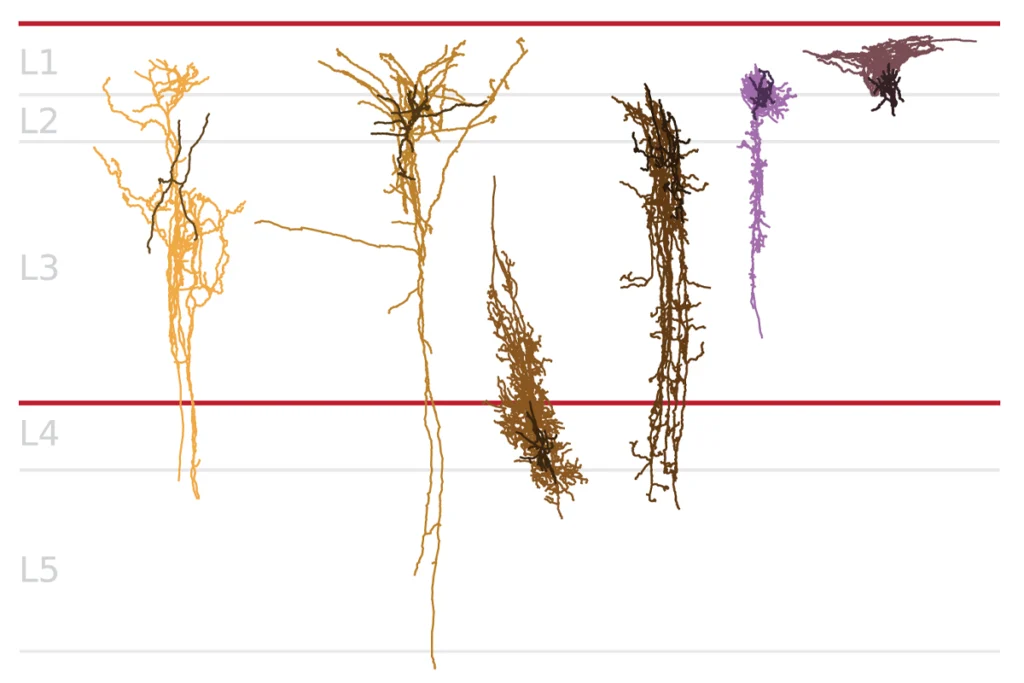
Scan of ‘missense’ mutations marks new suspects for autism risk
A large study of minute mutations in people with developmental conditions, including autism, has uncovered 200 potential risk genes.
A large study of minute mutations in people with developmental conditions, including autism, has uncovered 200 potential risk genes1.
The probe involved so-called ‘missense’ mutations, in which a swap of a single DNA letter alters one amino acid in a protein. Many of the newly identified genes perform familiar functions — regulating gene expression, neuronal development and neuronal junctions, or synapses.
Once confirmed, the findings open the door to a new group of genes associated with autism. “We could identify an entire new set of patients in terms of the genetic basis of autism,” says lead researcher Evan Eichler, professor of genome sciences at the University of Washington. The genes could also yield treatment targets, Eichler says.
Missense mutations often are harmless or produce only subtle effects, and are nearly as common in the general population as among people with autism. As a result, the harmful ones have been difficult to spot.
Eichler and his colleagues identified potentially harmful missense mutations using a three-pronged approach. They assessed the number of people who have the mutations, predicted the mutations’ severity with software, and looked for ones that cluster on a single gene.
“The new results are very reassuring that these three features in combination can be used to identify disease-causing mutations and pinpoint disease risk genes,” says Joseph Gogos, a neuroscientist at the Zuckerman Mind Brain Behavior Institute at Columbia University, who was not involved in the study.
Defining features:
Eichler and his team first analyzed non-inherited, or de novo, missense mutations in 8,477 people with autism, intellectual disability or developmental delay and 2,178 unaffected siblings and other unaffected individuals. The researchers collected the data from 24 published studies.
They found that people with any of these three conditions are 12 percent more likely to carry a missense mutation than are their siblings. The missense mutations in people with these conditions are also recurrent, up to three times as likely as those in controls to show up in more than one person.
What’s more, the most damaging mutations are between 20 and 80 percent more likely to be in an affected individual than in an unaffected one. The study appeared 19 June in Nature Neuroscience.
The findings suggest people with one of the developmental conditions have more recurrent missense mutations as well as more damaging ones than controls do.
The team then identified 35 genes that have significantly more missense mutations than expected in people with autism, intellectual disability or developmental delay. (By contrast, missense mutations occur more often than expected in only two genes in controls.) Of the 35 genes, 17 — including GRIN2B, PTEN and SCN2A — are known autism genes.
Lightning strike:
The researchers homed in on 40 sites within the genes at which missense mutations score high on their likely severity and occur in more than one person. They looked up 20 of those sites in a larger group of 17,688 people with autism or developmental delay from various genetic databases.
They found 21 new people with missense mutations at the same sites, boosting the evidence for these mutations being pathogenic. They did not find any of the mutations at these sites in a control group of approximately 3,000 people.
One frequently mutated site occurred in a gene called GRIA1. Five people with autism and one with intellectual disability carry this mutation, which is not found in the general population. “This is like lightning striking six times independently at the exact same point,” Eichler says.
GRIA1 encodes a receptor for a chemical messenger called glutamate, which boosts neural activity. It also plays a key role in early synapse development. A mutation in this gene seems to alter the protein’s ability to serve as a glutamate receptor, Eichler says.
Making sense:
To find other risk genes, the researchers took a new approach: Instead of flagging mutations that appear in more than one individual, they looked for those that land near each other within the same gene.
Their analysis pointed to 200 genes. The researchers found significantly more clusters of missense mutations in these genes among people with one of the conditions than among controls. What’s more, the clusters fall into known functional segments of a gene.
“To me, this is super exciting because it says not only we have identified a potential gene, but we know what part of the gene is, in fact, relevant,” Eichler says. Researchers might eventually target drugs to these segments, he says.
Knowing the location of missense mutations can help scientists understand their consequences and separate the harmless from the harmful ones, says Kaitlin Samocha, a postdoctoral fellow in Matthew Hurles’ lab at the Wellcome Trust Sanger Institute who was not involved with the new work. “Methods that search for clusters of mutations or other position-dependent effects are the way forward in this space,” she says.
Whether the 200 genes are truly linked to autism remains to be seen, but the findings open the door to a new set of genes for researchers to investigate. “We now have a hook to really discover the pathogenic missense genes,” Eichler says.
The team plans to examine the highlighted hotspots in all 200 genes in tens of thousands of people with one of the conditions.
References:
- Geisheker M.R. et al. Nat. Neurosci. 20, 1043-1051 (2017) PubMed
Corrections
This article has been modified to correct the name of the journal that published the study. It was Nature Neuroscience, not Nature Genetics.
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question