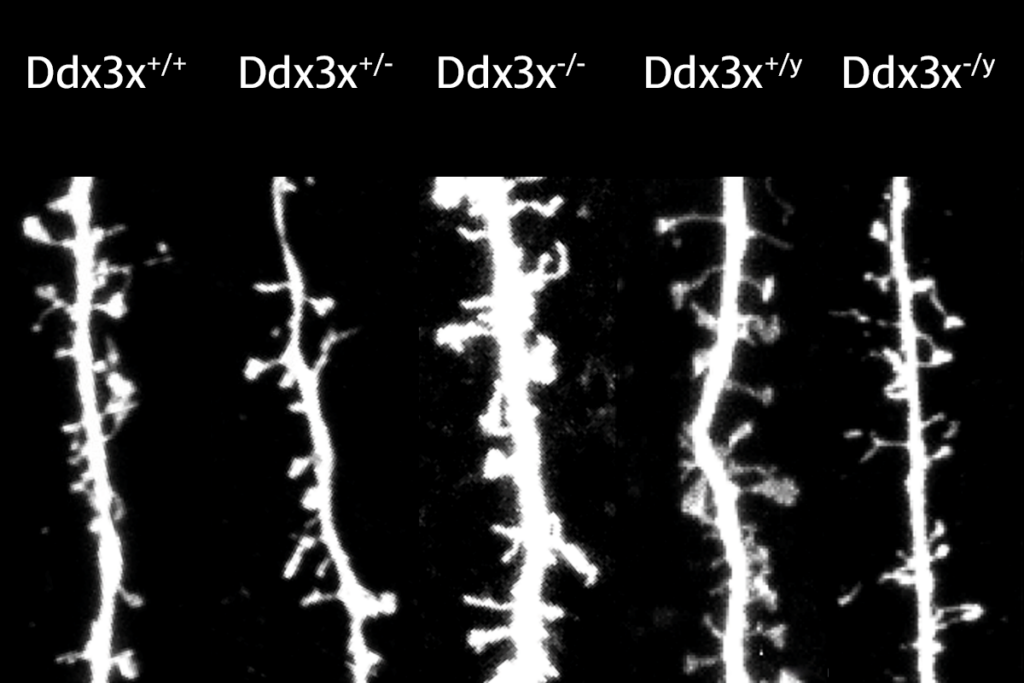
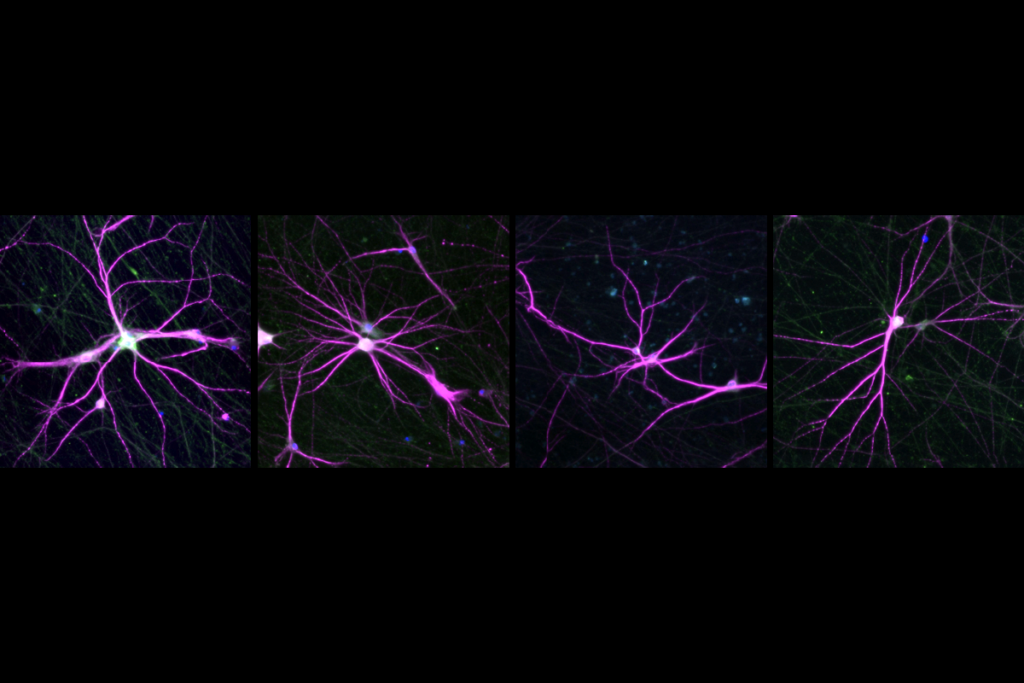
Two of the largest analyses of DNA sequences from people with autism have uncovered genes tied to the condition through a range of variant types. The work, published in two studies today in Nature Genetics, provides a more comprehensive picture of the condition’s genetic architecture and hints at how it may differ from that of other neurodevelopmental conditions.
Scientists typically hunt for de novo variants in sequences to spot autism-linked genes — an approach that has yielded roughly 100 genes. But alterations in these genes tend to have global effects on cognition and brain function, and people who carry them often represent only one portion of the autism spectrum.
“I think we’ve been biased in terms of the genes that we’ve identified,” says Wendy Chung, professor of pediatrics and medicine at Columbia University, who co-led one of the studies. To better understand the biology underlying brain function, behavior and autism, researchers have to look at genetic variation across the spectrum, she says.
Besides de novo variants, several other types — such as inherited variants and large duplications or deletions of DNA known as copy number variants (CNVs) — can contribute to autism. But insufficient sample sizes have limited researchers’ ability to detect genes strongly tied to the condition through these types of alterations.
The two new studies come from international collaborations that have amassed troves of genetic data. One analysis considers both de novo and inherited variants to uncover five new autism-linked genes, four of which are tied to a lower likelihood of cognitive impairment than five well-established autism-linked genes. The other study integrates data about single-letter changes in DNA, small insertions or deletions called indels, and CNVs to detect 373 genes linked to autism and other neurodevelopmental conditions, as well as 36 genes that show heightened variation in autism.
Together the studies expand the list of genes broadly tied to neurodevelopment, which researchers can use to build on previous work that aims to uncover how genes relate to specific conditions or traits, says Varun Warrier, a postdoctoral researcher in Simon Baron-Cohen’s lab at the University of Cambridge in the United Kingdom, who was not involved in the research.
“We don’t quite know what these genes are really associated with,” Warrier says, but the work points toward differences that may help researchers unravel heterogeneity among neurodevelopmental conditions.
C
hung and her colleagues analyzed genetic sequences from more than 42,607 people with autism and 79,670 of their parents and siblings. The team drew on previously published data from three projects — the Autism Sequencing Consortium, MSSNG and the Simons Simplex Collection — as well as new sequences from 35,130 people collected through an online registry called SPARK. (Like Spectrum, the Simons Simplex Collection and SPARK are both funded by the Simons Foundation Autism Research Initiative, where Chung is also director of clinical research.)The researchers used a two-stage approach: First they looked for genes enriched with de novo alterations. They also defined 25 gene sets based on existing genetic or functional evidence relevant to autism, such as genes expressed in neurons or those predicted by machine-learning tools to be tied to the condition. They then identified genes harboring variants that autistic people inherit more often than expected by chance within gene sets that are enriched with rare, inherited loss-of-function variants.
Overall, autism showed ties to de novo variants in 159 genes, and rare, inherited variants in another 260 genes. Known autism- and neurodevelopment-linked genes account for about 70 percent of the autism odds attributed to de novo variants, the team found, whereas these genes explain less than 20 percent of the likelihood conferred by rare, inherited variants.
“That means there is great value in identifying new risk genes through rare, inherited variants,” says Yufeng Shen, associate professor of systems biology at Columbia University, who co-led the work.
Another round of statistical tests incorporated an additional 8,116 samples from autistic children and their parents, again searching for genes enriched with de novo variants or harboring alterations that are over-inherited in autism among 367 genes on non-sex chromosomes. The analysis also compared rates of loss-of-function variants in 15,780 autistic people and 236,000 population controls.
The approach uncovered 60 genes strongly linked to autism, including 5 new ones: ITSN1, SCAF1, HNRNPUL2, MARK2 and NAV3. The first three genes in this list are tied to autism through multiple variant types. NAV3, however, is primarily linked to the condition through rare, inherited variants, the researchers report.

All of the novel genes but MARK2 have modest effects, the researchers found, and autistic people who carry variants in them are less likely to have cognitive impairment than people who harbor variants in a comparison set of five well-established autism-linked genes: CHD8, SCN2A, ADNP, FOXP1 and SHANK3. The genes with modest effects may be tied more closely to core autism traits than to brain development, the researchers say.
Although assessing cognition in people with autism can be challenging, the study is a step toward better reflecting the condition’s complexity, says Thomas Bourgeron, professor of genetics at the Institut Pasteur in Paris, France, who was not involved in the research. Taking cognitive impairment into account provides more nuance than just comparing autistic people with controls. “It’s a bit more precise,” he says.
I
n the other study, a team co-led by Michael Talkowski, director of the Center for Genomic Medicine at Massachusetts General Hospital in Boston, searched for single-letter changes in DNA and for indels in exome sequences from 63,237 people, 20,627 of whom have autism.They also used a method developed several years ago to detect CNVs in the exome data — drawn from a variety of repositories, including the Autism Sequencing Consortium, the Simons Simplex Collection, iPSYCH and SPARK — and calibrated the approach using whole-genome sequences from 7,165 people.
In total, that method yielded 17,774 rare, inherited and 662 de novo CNVs spanning three or more exons. The team used a statistical framework called TADA to combine the information about the CNVs, indels and single-letter DNA changes with a score that gauges a gene’s tolerance to variants that alter its function. That process revealed 72 autism-associated genes — a meaningful increase from a previous study that found 32 genes tied to the condition at the same level of statistical significance, the researchers reported.
Another analysis that combined sequences from the people with autism and 91,605 people from the Deciphering Developmental Disorders study — 31,058 of whom have a developmental condition — detected 373 genes tied to neurodevelopment. Genes identified among people with autism and among those with other developmental conditions overlapped, with 70 to 90 percent of genes highlighted in both groups.
Assessing variation among 464 genes associated with either condition through de novo mutations that likely affect gene function revealed 36 genes that harbor more variation among people with autism than they do in people with other developmental conditions. Another 82 genes are more commonly altered in people with developmental conditions.
These results provide a glimpse of some of the genes that might contribute to autism versus other developmental conditions, Talkowski says. The expression of genes tied to other developmental conditions is enriched in early neurodevelopmental cell types, according to an analysis of single-cell gene-expression data from the developing brain. Genes predominantly associated with autism, in contrast, are largely expressed in maturing neurons.
The findings are interesting, Warrier says, but children with other developmental conditions may have undiagnosed or co-occurring autism, and vice versa. “These are not watertight compartments,” he says, and better characterizing people in each group may help link genes to distinct traits, which is an ongoing challenge in the field.
Talkowski, too, notes that the two conditions are difficult to untangle. Which genes are tied to autism versus other neurodevelopmental conditions may shift as research continues, he says. Nonetheless, as more data become available, the study provides a framework for better understanding how genes and variant classes contribute to diverse neurodevelopmental conditions and traits.