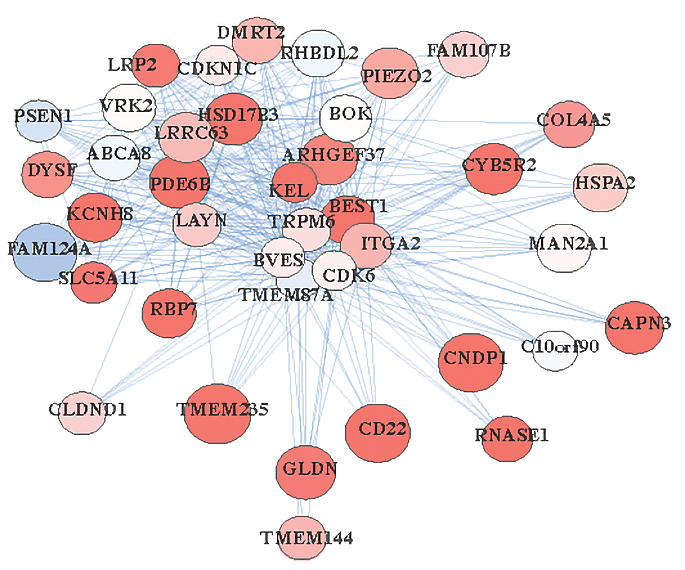
Differing gene functions across species may pose problems for autism models
Misaligned gene expression maps suggest that some autism-linked genes play distinct roles in mouse and human brains.
Some autism-linked genes, such as SHANK3 and SCN2A, may have different functions in mice and humans, according to a new study. The research identifies evolutionary changes in which genes are expressed together in the brain and raises new concerns about the limits of model organisms for studying neuropsychiatric conditions.
Mapping gene expression networks offers insight into genes’ functions, previous work has shown. Different cell types, for example, can be identified by the patterns of genes they tend to express — much as someone’s social network can suggest their roles in society, says lead investigator Daniel Geschwind, distinguished professor of neurology, psychiatry and human genetics at the University of California, Los Angeles.
In the new study, Geschwind and his colleagues examined how these networks vary across humans, non-human primates, mice and cultured human cells.
Expression patterns linked to neurons tend to be conserved between mice and humans, the study shows. But expression patterns related to glia — support cells in the nervous system that include astrocytes, microglia and oligodendrocytes — are not. Instead, glial gene expression in people seems to better match that of cultured cells, monkeys or chimpanzees.
What’s more, 70 autism-associated genes, as well as genes linked to other neuropsychiatric conditions, have divergent patterns of expression between humans and mice, the mapping revealed.
Researchers should not forgo studying these genes in mice, Geschwind says. “But making conclusions about neuropsychiatric conditions based on a mouse model in which the gene is not well preserved [between the two species] could be problematic.”
Model citizens:
Geschwind and his colleagues analyzed data from 7,287 samples of postmortem adult brain tissue collected by the Genotype-Tissue Expression project and 6,667 samples of mouse brain tissue collected in 30 previous studies. They generated gene expression networks across 12 human brain regions and 7 mouse brain regions and compared them.
Many gene networks are conserved, the team found. But the expression of 5,473 genes differs between the two species. Regions of cerebral cortex, the brain’s outer layer, showed the most significant differences, whereas patterns of expression in the cerebellum, which coordinates movement and timing, were more similar between the species.
Genes associated with glia showed more distinct patterns between people and mice, suggesting that evolution has driven more changes among these cells, Geschwind says. Genes linked to neurons, on the other hand, were more closely aligned. The findings were published in January in Genome Biology.
The researchers also generated gene expression networks for 2,933 tissue samples from six brain regions in macaque monkeys, baboons and chimpanzees. As with the mice, the non-human primate gene networks associated with neurons were similar to those seen in human brains. Expression patterns linked to other cell types more closely resembled those seen in humans than those from mice did, but those findings are not statistically significant.
Different parallels emerged for human brain organoids, clusters of cultured neural tissue grown from stem cells. Patterns of gene expression from eight previous organoid studies lined up with human gene expression for astrocytes and neurons, but not microglia or oligodendrocytes, the team found.
Knowing limits:
The organoid findings make sense, says Alysson Muotri, professor of cellular and molecular medicine at the University of California, San Diego, who was not involved in the work. For instance, most brain organoid models do not naturally contain microglia, because those cells emerge from a different cellular source than neurons do, he says. The new work suggests that adding microglia to organoids could make them a better model for autism, Muotri says.
That is the kind of guidance researchers should take away from the work, Geschwind says. Rather than crowning the “best” overall model organism, the results “highlight the pros and cons” of each for a given gene, he says.
As a researcher gearing up for new work on a mouse model of autism, the findings actually inspire confidence in what the model can and cannot do well, says Santhosh Girirajan, associate professor of genomics at Pennsylvania State University in University Park, who was not involved in the study.
“If you don’t keep the limitations [of a model system] in your mind, your interpretation of what you find may not be accurate,” he says.
One limit on the utility of the new results, particularly for autism research, is that the study uses only adult brain tissue, which may not accurately represent gene expression during development. Geschwind and his colleagues plan to expand their work to include prenatal and postnatal networks when the data for those periods become available.
Explore more from The Transmitter

Exon-skipping approach boosts levels of key Rett syndrome protein