Networks of genes altered in autism brains, study says
Two networks of genes are abnormally expressed in the brains of people with autism, according to a study published today in Nature.
-
Two networks of genes are abnormally expressed in the brains of people with autism, according to a study published today in Nature1.
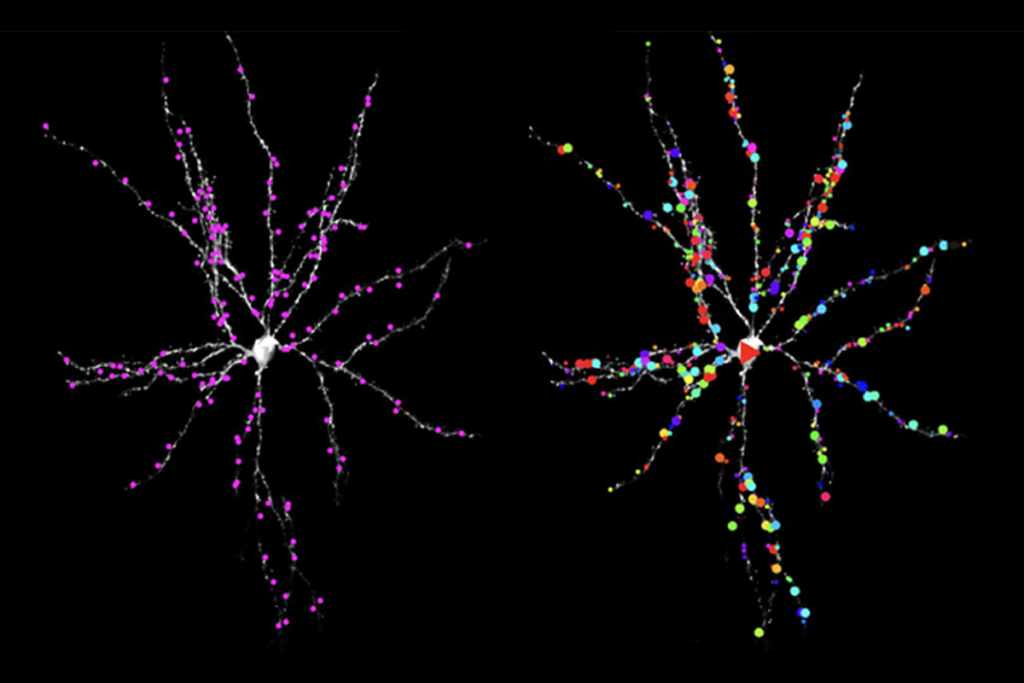
Analyzing postmortem brain tissue, the researchers discovered that, compared with controls, genes involved in cell connectivity tend to be expressed at lower levels in autism brains, and genes related to immune cells at higher levels.
Autism is known for its diversity in symptoms and in the genes that might cause it. “[But] there is a remarkable consistency in the molecular changes that are occurring in the brain,” notes lead investigator Daniel Geschwind, distinguished professor of neurology, psychiatry and human genetics at the University of California, Los Angeles.
The study turned up several unexpected findings. For example, the frontal lobe and temporal lobe in healthy controls show significant differences in gene expression, reflecting their distinct cell types and functions. But in the autism brains, “those differences are essentially wiped out,” Geschwind says. Many of these genes are first turned on during embryonic development, he says, suggesting that the abnormal trajectory of autism brains begins early.
“This has never been reported before — it’s definitely an original contribution and an advance,” notes John Allman, professor of neurobiology at the California Institute of Technology, who was not involved in the study.
Another big finding: A2BP1, one of the genes expressed at low levels in autism brains, leads to the abnormal transcription of a host of other genes in autism brains — potentially creating problems that contribute to autism. “This could be where the real action is happening,” Allman says.
Marking modules
In the past few years, researchers have zoomed in on the genetic underpinnings of autism and other developmental disorders. But work on the next biological step — the complex ways in which DNA turns into RNA and then into protein — is rare.
A few years ago, Geschwind’s team identified about two dozen networks of genes that tend to transcribe DNA into RNA at the same time2. Each network, or ‘module,’ holds genes involved in similar tasks, such as those that mark particular cell types, or that maintain neuronal connections related to the chemical messenger glutamate.
In the new study, the researchers looked for similar networks in postmortem brain tissue from 19 individuals with autism and 17 controls from tissue banks overseen by the Autism Tissue Program. They focused on tissue from the cerebral cortex, the outer layer of the brain that’s important for planning and learning.
They identified 18 modules of genes expressed together in the cortex, some of which change with the age or sex of the samples. Two modules are significantly different in the autism group compared with controls, they found.
The first, dubbed M12, comprises 209 genes expressed at lower levels in the brains of people with autism than in controls. It includes genes related to the synapse — the junction between neurons — as well as those involved in the branching of neurons and the movement of neurotransmitters.
The second module, M16, holds 235 genes that are expressed at higher levels in autism brains compared with controls. The list includes genes related to star-shaped cells called astrocytes, and activated microglia — immune cells that have been implicated in autism and other psychiatric disorders.
These findings all held up in a replication sample of nine autism brains and five controls.
The researchers also honed in on one of the M12 genes, A2BP1, which had been fingered in a couple of autism studies.
Stretches of DNA can be transcribed, or ‘spliced,’ in different ways, which can have significant effects on the function of the protein. A2BP1 helps cells edit RNA correctly. By deeply sequencing RNA, the study found that in autism brains, A2BP1 is expressed at lower levels than normal, altering the editing of several other genes. One of its targets, notably, is GRIN1, a gene that encodes part of the ubiquitous NMDA receptor. This protein is crucial for learning, which is often impaired in individuals with autism. “That’s a home run,” Allman says.
Developmental questions
Identifying these common molecular pathways bodes well for therapeutic approaches to autism, notes Jon McClellan, professor of psychiatry and behavioral sciences at the University of Washington in Seattle.
“The goal of all of this is to try to get more specific treatment,” McClellan says. “If you figure out some critical neurobiological pathway, then the hope is that it won’t matter what the particular cause is, the treatments correcting that pathway will still work.”
Still, it’s difficult to discern whether the hundreds of gene expression differences are signatures of autism’s cause, or the consequence of living with the disorder. “It’s like any big complicated system. If something happens early on, the downstream effects can ripple everywhere,” McClellan says.
To get at this timing question, the researchers looked at which of the abnormally expressed genes have been identified in genetic studies of autism. They found that the M12 module is significantly enriched with autism candidate genes — such as APBA2, CNTNAP2 and SLC25A12 — whereas the M16 module is not.
This suggests that the M12 expression differences stem from variations in inherited DNA, whereas those in M16 are caused by changes that crop up after birth. Geschwind says he suspects that inherited problems in synapse-related genes cause the brain to compensate over time, leading to the changes in immune-related genes. In fact, the brains of people with schizophrenia and Alzheimer’s disease also show higher expression of immune genes.
Not every autism sample showed differences in M12 and M16, he notes, but those with dampened M12 expression also show higher expression of M16 genes. “This suggests that the two processes are connected,” he says.
Other researchers note, however, that this approach doesn’t necessarily capture changes in early development. The tissue used in this study was taken from individuals ranging from 2 to 81 years old, but the vast majority came from adults.
“We know autism is a developmental disorder, so we need to study the developing brain,” notes Manuel Graeber, professor at the Brain and Mind Research Institute of the University of Sydney in Australia. But postmortem tissue is a scarce resource. “More than anything else, I think what this study demonstrates is the need for more and better brain banking,” Graeber says.
Geschwind’s team is analyzing the expression of autism candidate genes in neural stem cells taken from the brains of human fetuses. “That could provide another line of evidence for very early alterations in the brain,” Geschwind says.
References:
Explore more from The Transmitter