New algorithm detects DNA insertions and deletions
A new algorithm accurately detects large DNA insertions and deletions in the protein-coding regions of the genome.
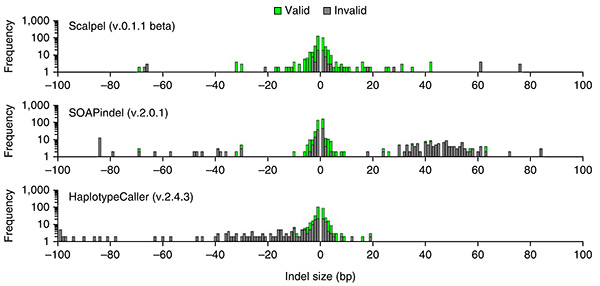
Accurate assessment: Scalpel (top) is more likely to correctly identify DNA insertions and deletions (green) than are older methods.
A new algorithm accurately detects large DNA insertions and deletions, together referred to as ‘indels,’ in the exome, or protein-coding regions of the genome, according to a study published 17 August in Nature Methods1.
Compared with their unaffected siblings, children with autism have been found to have almost twice as many spontaneous indels that are likely to disrupt protein production2. The researchers confirmed this pattern with their new method, called Scalpel. The method showed that these sorts of mutations in genes involved in neural development are particularly common in people with autism.
Other methods to detect indels typically involve sequencing long strands of DNA, and are expensive and slow. Researchers then compare DNA sequences with the reference genome, assembled from the sequences of multiple healthy people. Cheaper methods, which involve chopping DNA into tiny pieces, are effective at detecting small mutations but tend to miss larger ones.
Scalpel provides algorithms for piecing together short DNA sequences that are part of the same gene. This yields longer sequences, which are easier to match to the reference genome but less expensive to produce than long sequences.
The researchers tested Scalpel and eight other algorithms used for detecting indels to analyze simulated DNA sequences. Scalpel and two others performed best in this step. The researchers then used those three methods to analyze parts of a real genome. They checked the results by sequencing small portions of DNA using a slow and expensive method.
They found that Scalpel detects indels longer than 30 base pairs more accurately than the other methods do.It also excels at detecting these mutations in regions that already have repetitive DNA sequences.
The researchers used Scalpel to analyze genetic data from 593 people with autism and their families, taken from the Simons Simplex Collection. This database houses genetic and other data on families that have one child with autism and unaffected siblings and parents. (The collection is funded by the Simons Foundation, SFARI.org’s parent organization.)
Scalpel detected 3.3 million indels in the families. Children with autism and unaffected siblings all have indels, but the researchers found 35 indels that would likely affect proteins in children with autism, compared with 16 in unaffected siblings.
Of the 35 found in children with autism, 8 are in targets of FMRP, the fragile X syndrome protein.
References:
1. Narzisi G. et al. Nat. Methods Epub ahead of print (2014) PubMed
2. Iossifov I. et al. Neuron 74, 285-299 (2012) PubMed
Recommended reading

Building an autism research registry: Q&A with Tony Charman
CNTNAP2 variants; trait trajectories; sensory reactivity
Brain organoid size matches intensity of social problems in autistic people
Explore more from The Transmitter
Cerebellar circuit may convert expected pain relief into real thing