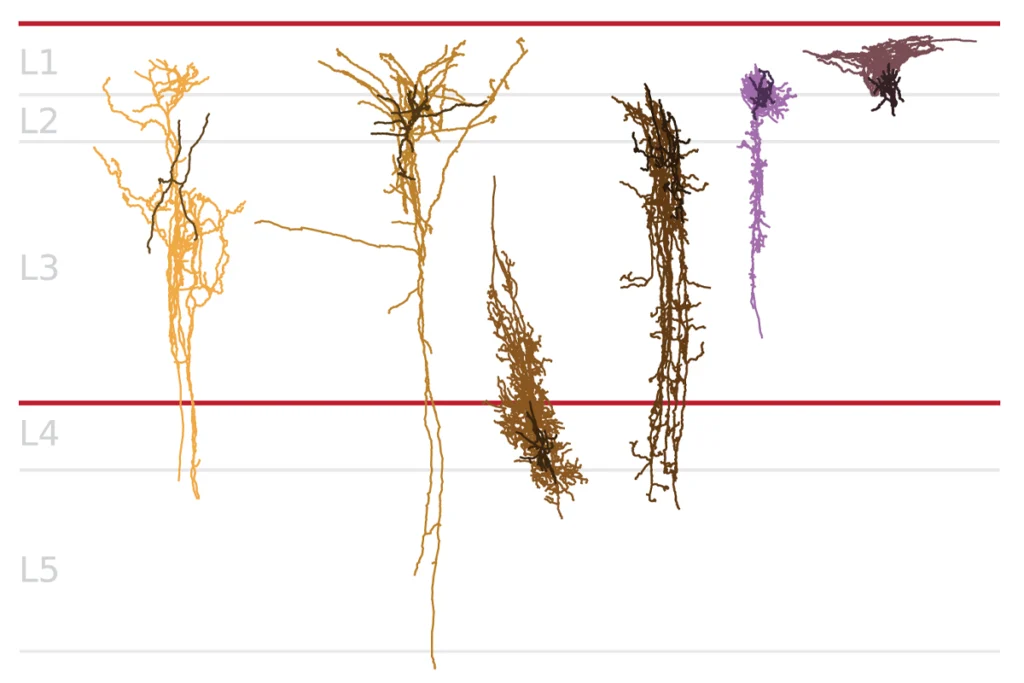
Rho family of enzymes at crossroads of autism
A number of autism risk factors converge on one cellular pathway: abnormal remodeling of the cell’s structural systems through the signaling protein Rho, says SFARI’s associate director for research, Alan Packer.
Over the past few years, the projected number of risk factors for autism has reached between 500 and 1,000 genes. More recently, the field has begun to identify points of convergence at which multiple genes affect the same cellular functions1.
Fortunately, insights from analyses of known risk genes give us reason to think that the list of candidate genes will advance the field substantially. There are some emerging common themes, including signaling pathways that strengthen neuronal connections in response to experience2, 3 and the collection of targets of FMRP, the protein missing in fragile X syndrome4.
Variations in common single-nucleotide polymorphisms5 and in patterns of gene expression6, each of which has at least some predictive power in diagnosing autism, support the role of these critical pathways in autism.
All of the genetic heterogeneity we’re encountering has led to some concern in the field that these genes might lead us in 500 different directions. But in my humble opinion, with just an Internet connection and a few hours to spare, it is possible to come up with interesting hypotheses as to how some of these genes converge on a relatively small number of cellular processes.
In this column, I discuss one such point of convergence: abnormal remodeling of the actin cytoskeleton, the cell’s structural support network, during the growth of axons and dendrites — the long projections and signal-receiving branches of neurons, respectively. A specific focus will be on the role of the Rho family of GTPases, which are known to have key roles in regulating actin dynamics.
Actin remodeling:
Several studies, particularly surveys of copy number variants (CNVs) — large deletions or duplications of DNA — associated with autism risk, have highlighted this process of actin remodeling7, 8. But new data from efforts to sequence the coding region of the genome, or exome, have confirmed and extended its importance.
In addition, while I was writing this article, a new study appeared to reach similar conclusions based on a more global, unbiased survey of the literature9.
The place to start is with a study published in January and summarized here, by Jocelyn Krey, Ricardo Dolmetsch and their colleagues. The study uses rodent and human induced pluripotent stem (iPS) cells to study the mutation that causes Timothy syndrome10. This mutation is in a subunit of a calcium channel encoded by the CACNA1C gene, and leads to a complex syndrome frequently accompanied by autism.
The new study shows that the Timothy syndrome mutation promotes a net reduction in dendrite length, at least in all of the cells carrying the CACNA1C mutation that the researchers examined.
Through an additional series of elegant experiments, the researchers show that the mutant calcium channel affects dendrite dynamics by activating a series of small GTPases, a family of enzymes with a range of cellular functions.
This cascade culminates in the hyperactivation of one such GTPase called RhoA (one of several Rho family members), and expression of a permanently active form of RhoA in typical neurons leads to the dendrite retraction seen in cells carrying the Timothy mutation. This is a new and important molecular insight into the mechanism underlying the syndrome.
Rho signaling comprises a well-studied group of pathways that regulate remodeling of the actin cytoskeleton, which takes place in all cells that need to move or change shape. One arm of this signaling pathway begins with RAS GTPase and activates other molecules, such as CDC42, N-WASP, PAK and LIMK1.
Another arm of the pathway signals through RhoA, activating molecules such as ROCK and also LIMK1. LIMK1 is a kinase that adds a phosphate to, and inhibits, a protein called cofilin. Cofilin helps disassemble the actin network that lends the cell its shape. For that shape to change, or to allow cell movement, the actin network needs to disassemble and then reassemble.
Skeleton crew:
Studies have implicated Rho-dependent regulation of the actin cytoskeleton in autism. A survey of CNVs in individuals from the Autism Genome Project suggests that deletions or duplications of genes in the Rho pathway are more common in individuals who have autism or intellectual disability than in controls7.
A 2011 study reinforced this finding with an analysis of CNVs in the Simons Simplex Collection, which includes families that have only one child with autism8. (The collection is sponsored by the Simons Foundation, SFARI.org’s parent organization.) The researchers discussed ways in which dysregulation of the Rho pathway might explain the underlying neuronal pathology of autism.
As powerful as those studies are, one limitation in interpreting them lies in the fact that few of the CNVs are both small and common enough to implicate individual genes with strong statistical significance. It could be argued that studies such as the iPS study of Timothy syndrome, which focus on individual risk genes that lead to syndromic autism, address this gap.
I’ve already mentioned that LIMK1 responds to RAS and RhoA signaling. Importantly, LIM1K is at chromosome 7q11.23, and is deleted in Williams syndrome, a disorder characterized in part by hypersociability. Duplication of this region, on the other hand, is strongly associated with autism11.
Elevated RhoA activity seen in Timothy syndrome would be expected to boost LIMK1 activity as well. Because of this, the association of both Timothy syndrome and 7q11.23 duplication with autism is consistent with elevated RhoA-dependent signaling playing a role in autism.
Researchers are still working on a 7q11.23 duplication mouse, but a mouse lacking LIMK1 has elongated dendritic spines. This is also consistent with what one would expect from dampened RhoA/LIMK1 signaling.
Excitingly, exome-sequencing efforts in autism have also identified a number of autism risk genes that are linked to RhoA signaling and the cell biology of cytoskeletal remodeling. One example is EPHB2, which encodes one of the ephrin receptors known to be involved in axon guidance and cell migration, among other things.
A 2010 study showed that EPHB2 adds a chemical phosphate to a molecule called Ephexin5, and that this phosphorylation enables UBE3A — a ubiquitin ligase linked to autism — to degrade Ephexin512. Ephexin5 also activates RhoA. So, mutation of EPHB2 (as in some cases of autism) would lead to elevated Ephexin5, which would promote RhoA activation. Hyperactivation of RhoA or RhoA effectors is, again, consistent with what one would expect from Timothy syndrome and 7q11.23 duplication.
Another likely autism risk gene is DYRK1A, a kinase encoded by a gene that is in the Down syndrome critical region on chromosome 21. DYRK1A has many roles in brain development, but a study last year showed that one of its functions is to regulate the actin cytoskeleton by phosphorylating N-WASP — a protein that plays a role in actin polymerization13.
Finally, studies have implicated spontaneous mutations in CUL3 in autism susceptibility. CUL3 is a member of the cullin family of scaffold proteins that enable the assembly of ubiquitin ligase complexes, which degrade proteins.
Complex effects:
In 2009, before this gene was on the radar of anyone studying autism, researchers showed that CUL3 is part of a complex that ubiquitinates and degrades RhoA14. Dampening CUL3 expression prevents the degradation of RhoA and results in the production of abnormal actin stress fibers — structures that are required for cell contraction and movement.
The researchers also identified a family of RhoA-binding adaptor molecules that allow CUL3-containing complexes to ubiquitinate RhoA1. These adaptors go by the unlovely acronym ‘BACURDs’ (BTB-containing adaptor for CUL3-mediated RhoA degradation). The reader can be forgiven for thinking that BACURDs don’t have anything to do with autism, as it is not widely known that another name for BACURD1 is KCTD13.
KCTD13 is in the 16p11.2 region that is associated with autism and other neurodevelopmental disorders. KCTD13 is one of the best candidates for a gene in this region that contributes to the features of autism15.
The absence of either KCTD13 or CUL3 is likely to lead to hyperactivation of the RhoA pathway. This association is a particularly satisfying connection, linking a recurrent single-gene mutation and a recurrent CNV to the same pathway.
CUL3 itself should receive a great deal more attention from the autism research community, as it has also been implicated in the regulation of sleep16 and control of gene expression17.
What is the consequence of dysregulation of Rho GTPases and the actin cytoskeleton? Much of the work to date has suggested that altered dendritic spine dynamics lead to changes in synaptic plasticity — how brain signals adapt to experience. For example, mice lacking LIMK1 have elongated spines and enhanced long-term potentiation, a process that underlies synaptic plasticity18.
One might assume that any mutation boosting LIMK1 expression or activity would have the opposite effect. However, the effects of altered Rho-dependent signal transduction on dendrite and spine morphology and on plasticity are not always easy to predict19. They may depend on the protein in question and whether its alteration is transient or sustained.
Additional possibilities are raised by a recent study that showed that a SYNGAP1 mutation, which leads to inhibition of cofilin activity (SYNGAP1 is a negative regulator of RAS), leads to more rapid maturation of dendritic spine synapses20. A major challenge to the field will be to link these changes in dendrite and spine morphology to the function of key neural circuits involved in autism.
All these links suggest many experiments. To begin with, we don’t yet know if mutations in the genes mentioned above will have the expected effects on Rho-dependent signaling, dendrite and spine development, and on synaptic plasticity and behavior. These predictions need to be rigorously tested in mouse models.
If the results are as expected, we can begin to think about generating mice with more than one mutation, with an eye toward rescuing some of the cellular, and perhaps even behavioral, symptoms. For example, one might expect that a mouse carrying the Timothy syndrome mutation would have fewer symptoms if it also had only one copy of LIMK1.
By the same token, a CUL3 mutation might exacerbate symptoms in a 16p11.2 deletion mouse, as both CUL3 and KCTD13 degrade RhoA. Also, researchers can test small-molecule inhibitors of RhoA, such as Rhosin21.
We know that the functions of many proteins in the Rho pathway are multifaceted, and the various mutant phenotypes complex. But experiments like these should tell us how much can be attributed to Rho-dependent pathways, and how much might be corrected by targeting them.
Alan Packer is a senior scientist at The Simons Foundation Autism Research Initiative.
References:
1. State M.W. and N. Sestan Science 337, 1301-1303 (2012) PubMed
2. Zoghbi H.Y. and M.F. Bear Cold Spring Harb. Perspect. Biol. 4 (2012) PubMed
3. Ebert D.H. and M.E. Greenberg Nature 493, 327-337 (2013) PubMed
4. Iossifov I. et al. Neuron 74, 285-299 (2012) PubMed
5. Skafidas E. et al. Mol. Psychiatry Epub ahead of print (2012) PubMed
6. Kong S.W. et al. PLoS One 7, e49475 (2012) PubMed
7. Pinto D. et al. Nature 466, 368-372 (2010) PubMed
8. Gilman S.R. et al. Neuron 70, 898-907 (2011) PubMed
9. Zeidán-Chuliá F. et al. Neuromolecular Med. Epub ahead of print (2013) PubMed
10. Krey J.F. et al. Nat. Neurosci. 16, 201-209 (2012) PubMed
11. Sanders S.J. et al. Neuron 70, 863-885 (2011) PubMed
12. Margolis S.S. et al. Cell 143, 442-455 (2010) PubMed
13. Park J. et al. J. Cell Sci. 125, 67-80 (2012) PubMed
14. Chen Y. et al. Mol. Cell 35, 841-855 (2009) PubMed
15. Golzio C. et al. Nature 485, 363-367 (2012) PubMed
16. Pfeiffenberger C. and R. Allada PLoS Genet. 8, e1003003 (2012) PubMed
17. Yanagiya A. et al. Mol. Cell 46, 847-858 (2012) PubMed
18. Meng Y. et al. Neuron 35, 121-133 (2002) PubMed
19. Murakoshi H. and R. Yasuda Trends Neurosci. 35, 135-143 (2012) PubMed
20. Clement J.P. et al. Cell 151, 709-723 (2012) PubMed
21. Shang X. et al. Chem. Biol. 19, 699-710 (2012) PubMed
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question