Control centers for genes rife with autism-linked DNA blips
DNA sequences called enhancers — which boost the expression of genes from within or outside them — are enriched for genetic variants linked to autism, suggests a new study. The finding may help researchers understand how variants outside genes contribute to autism.

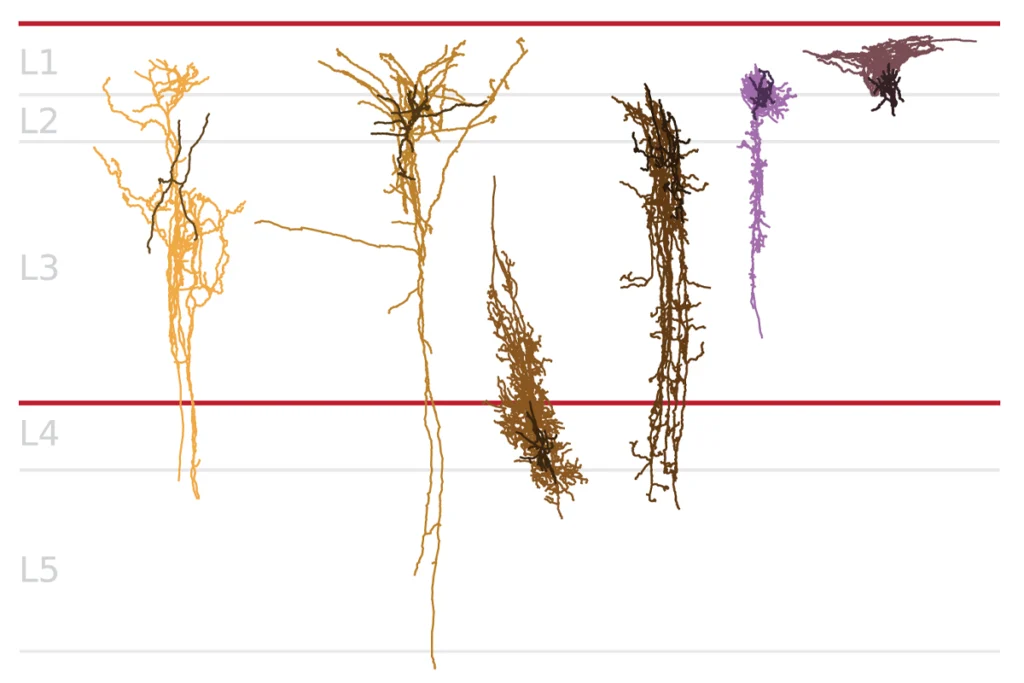
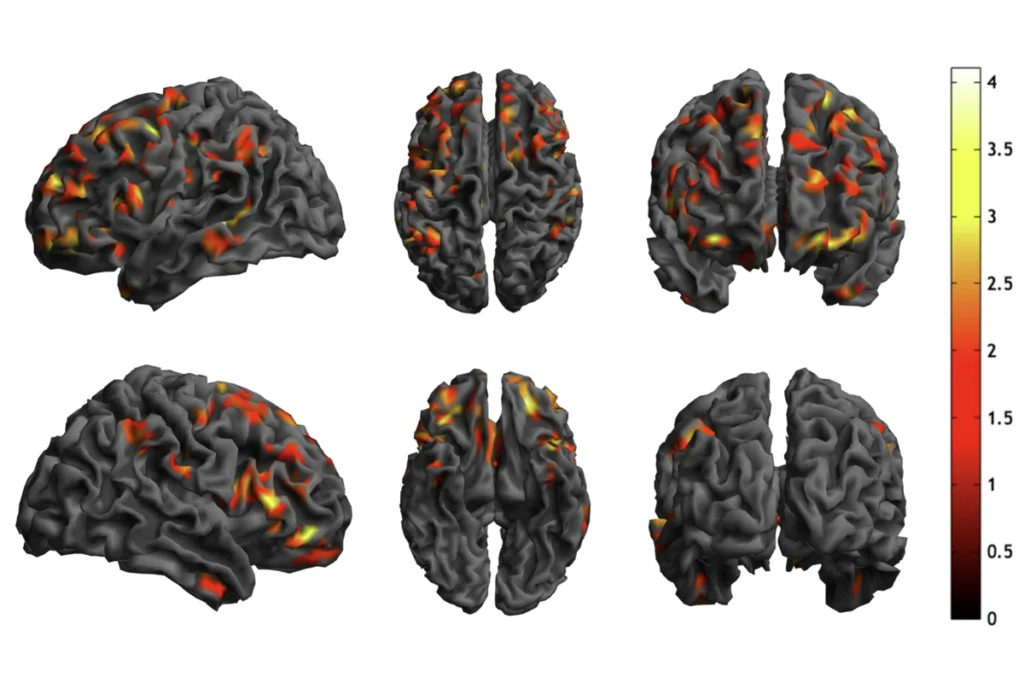
Partners in regulation: Enhancers and their potential target genes are often expressed together (red) — or repressed together (green) — depending on the brain region.
DNA sequences called enhancers — which boost the expression of genes from within or outside them — are enriched for genetic variants linked to autism, suggests a study published 13 July in Nature Neuroscience. The finding may help researchers understand how variants outside genes contribute to autism1.
Studies attribute at least half the risk for autism to genetics. So far, investigators have mostly tried to uncover genetic variants — differences in the DNA sequence — that disrupt genes. But these sequences, collectively known as the exome, constitute no more than 3 percent of the genome.
The remaining 97 percent contains enhancers and other elements that control genes, and remains relatively unexplored. The new paper suggests that these unstudied regions are rich with variants, known as single nucleotide polymorphisms (SNPs), that may play a role in disorders such as autism.
“[We need] to start looking more carefully into regulatory regions when we sequence the genomes of people with autism,” says lead investigator Irina Voineagu, senior lecturer of biological sciences at the University of New South Wales in Sydney, Australia.
The paper provides a shortlist of enhancer sequences for scientists to include in their studies of autism risk.
“Now we have a set of enhancers that we can look for variants within and potentially test their functional impacts,” says Dan Arking, associate professor of genetic medicine at Johns Hopkins University in Baltimore, Maryland, who was not involved in the study.
Noncoding boosters:
Last year, researchers cataloged more than 43,000 enhancers in the human genome2. These sequences generally display two telltale features: They tend to carry methyl groupson histones — the proteins associated with DNA — and they are made into molecules called enhancer RNAs (eRNA).
To pinpoint sequences relevant to brain disorders, Voineagu and her colleagues searched through the catalog and pulled out more than 900 that act in the brain. They then used postmortem brain tissue from four men who had no known disorders to find 314 enhancers that they could confirm become eRNAs, and so are likely to control genes.
To further whittle their list, the researchers identified enhancers that are enriched in the brain. They also looked for enhancers noted as carrying methyl marks in another massive dataset, the Roadmap Epigenomics Project3.
The collection is a valuable resource for autism research, says Nikolaos Mellios, assistant professor of neuroscience at the University of New Mexico, who was not involved in the study. “This paper should allow us to study eRNAs and enhancer regions to see why they are so important for the brain, and why they are linked to autism.”
To explore whether any of these enhancers carry genetic variants associated with neuropsychiatric disorders, Voineagu’s team combed through data from a study that identified more than 1 million SNPs shared by people with one of five neurological conditions: autism, attention deficit hyperactivity disorder (ADHD), depression, bipolar disorder and schizophrenia4.
They found that the SNPs within these enhancers are more strongly linked to autism than to any of the other four disorders. Still, even those are only weakly tied to autism, Arking says. To make the connections more definitive, he says, researchers will need to examine the sequences in tens of thousands of individuals with autism.
The new study offers a few more tantalizing hints of a link between enhancers and autism.
For example, Voineagu and her colleagues analyzed 401genes linked to autism to see how frequently the regions that control them contain an enhancer from the researchers’ shortlist. (They limited their analysis to regulatory regions within genes). They found that these regions carry enhancers more frequently than do genes tied to ADHD or schizophrenia.
Using data from the 2014 study, the researchers then identified 19 networks of genes and enhancers that are expressed at the same time in the same region of the brain.
One of these networks is highly expressed in the cerebellum, an autism-linked brain region that controls movement. This network contains 31 enhancers — nearly one-third of the researchers’ shortlist.
Three other networks are more active in fetal brain samples than in the brains of adults, suggesting that they play a role in brain development. These networks contain 12 enhancers, including one located within SHANK2 and another in BCL11A, two autism-linked genes. An additional enhancer in one of the fetal networks is highly expressed along with SOX4, FABP7 and AUTS2, three genes moderately linked to autism.
It’s not yet clear whether the variants in enhancers alter their function and whether the enhancers contribute to autism risk.
“It’s really difficult to know whether a single change in an enhancer will really affect it without actually testing that using some experimental system,” says Brian O’Roak, assistant professor of molecular and medical genetics at Oregon Health & Science University, who was not involved in the study.
References:
1. Yao P. et al. Nat. Neurosci. 18, 1168-1174 (2015) PubMed
2. Andersson R. et al. Nature 507, 455-461 (2014) PubMed
3. Zhu J. et al. Cell 152, 642-654 (2013) PubMed
4. Cross-disorder group of the Psychiatric Genomics Consortium Lancet 381, 1371-1379 (2013) PubMed
Recommended reading
Okur-Chung neurodevelopmental syndrome; excess CSF; autistic girls

New catalog charts familial ties from autism to 90 other conditions
Explore more from The Transmitter

Karen Adolph explains how we develop our ability to move through the world
Microglia’s pruning function called into question